Haplogroup I-M253
| Haplogroup I1 (M253) | |
|---|---|
| Possible time of origin | 3,170–4,600[1]–5,070 BP (today's diversification)[2][3] (previously 11,000 BP[4] to 33,000 BP[5]) 27,500 (diversification with I2-FGC77992)[1] |
| Possible place of origin | Northern Europe[6] |
| Ancestor | I* (M170) |
| Descendants | I1a (DF29/S438); I1b (S249/Z131); I1c (Y18119/Z17925) |
| Defining mutations | M253, M307.2/P203.2, M450/S109, P30, P40, L64, L75, L80, L81, L118, L121/S62, L123, L124/S64, L125/S65, L157.1, L186, L187 |
Haplogroup I-M253, also known as I1, is a Y chromosome haplogroup. The genetic markers confirmed as identifying I-M253 are the SNPs M253,M307.2/P203.2, M450/S109, P30, P40, L64, L75, L80, L81, L118, L121/S62, L123, L124/S64, L125/S65, L157.1, L186, and L187.[7] It is a primary branch of Haplogroup I-M170 (I*).
Haplogroup I1 is believed to have been present among Upper Paleolithic European hunter-gatherers as a minor lineage but due to its near-total absence in pre-Neolithic DNA samples it cannot have been very widespread. Neolithic I1 samples are very sparse as well, suggesting a rapid dispersion connected to a founder effect in the Nordic Bronze Age. Today it reaches its peak frequencies in Sweden (52 percent of males in Västra Götaland County) and western Finland (more than 50 percent in Satakunta province).[8] In terms of national averages, I-M253 is found in 38-39% of Swedish males,[9][10][8] 37% of Norwegian males,[11][12][13] 34.8% of Danish males,[14][15][16] 34.5% of Icelandic males,[17][18][19] and about 28% of Finnish males.[20] Bryan Sykes, in his 2006 book Blood of the Isles, gives the members – and the notional founding patriarch of I1 the name "Wodan".[21]
All known living members descend from a common ancestor 6 times younger than the common ancestor with I2.[1]
Before a reclassification in 2008,[22] the group was known as I1a, a name that has since been reassigned to a primary branch, haplogroup I-DF29. The other primary branches of I1 (M253) are I1b (S249/Z131) and I1c (Y18119/Z17925).
More than 99% of living men with I1 belong to the DF29 branch which is estimated to have emerged in 2400 BCE.[23][24] All DF29 men share a common ancestor born between 2500 and 2400 BCE.[25] The oldest ancient individual with I1-DF29 found is Oll009, a man from early Bronze Age Sweden.[26][27]
A 2024 study found that I1-M253 expanded rapidly during a migration from the eastern or northeastern parts of Scandinavia into Southern Sweden, Denmark and Norway around 2000 BC and was associated with the introduction of stone cist burials.[28] The study concluded that the spread of Y-DNA haplogroup I1 was associated with a genetic cluster labelled as "LNBA phase III" and that this genetic cluster formed the predominant ancestry source for Bronze Age, Iron Age and Viking Age Scandinavians, as well as other ancient European groups with a documented Scandinavian or Germanic association (for example, Anglo-Saxons and Goths; Extended Data Fig. 6e).
Origins
[edit]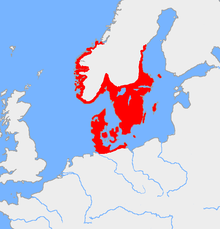
While haplogroup I1 most likely diverged from I* as early as 27,000 years ago in the Gravettian, so far DNA studies have only been able to locate it in three Paleolithic and Mesolithic hunter-gatherers. As of November 2022, only 6 ancient DNA samples from human remains dating to earlier than the Nordic Bronze Age have been assigned to haplogroup I1:
- BAL051. A hunter-gatherer from the Azilian in Spain in 11,466 BCE classified as having a now extinct branch of I-Z2699.[24][29]
- Burial SF11 Date: 7500 BC– A Scandinavian hunter-gatherer with the label SF11 found on Stora Karlsö on Gotland. SF11 was found to have carried 9 of the 312 SNPs that define haplogroup I1. SF11 was classified as I1-Z2699*.[30][31][32][33] SF11 was not assigned to a specific archaeological culture due to the skeleton being found in the Stora Förvar cave on Stora Karlsö.
- Burial BAB5: Assumed a date of 5600-4900 BC according to archaeological context, although without C14 dating – An individual sample from Balatonszemes-Bagodomb labelled BAB5, from Hungary.[34] BAB5 was found to have carried 1 of the 312 SNPs that define haplogroup I1. BAB5 may also be classified as I1-Z2699*.[35] The skeletal remains of BAB5 have not been radiocarbon dated, and the 2015 study provides no analysis of the sample's autosomal ancestry. A 2023 study with samples from the same site features several Migration Period Langobard aDNA samples, putting doubts to the archaeological dating of sample BAB5.[36]
- Burial RISE179 Date: 2010-1776 BC – An individual found in a kurgan burial dating to the late Neolithic Dagger Period in Scandinavia labelled RISE179.[37] The grave is located close to Abbekås on the south coast of Skåne RISE179 had a genetic affinity to the populations of the Corded Ware culture and the Unetice culture.[37]
- Burial oll009 Date:1930-1750 BC - A LNBA individual from Southern Sweden. oll009 and was sequenced in the study titled "The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon".[38] Oll009 is dated to the Scandinavian late Neolithic and was found in a burial in Sweden close to Öllsjö on the east coast of Skåne. Similar to RISE179, he carried a high percentage of Western Steppe-Herder ancestry and had a genetic affinity to the population of the Battle Axe culture and other populations of the Corded Ware horizon.[39] oll009 has Y11204 but does not seem to have Y164553 or Y11205.[40]
Despite the high frequency of haplogroup I1 in present-day Scandinavians, I1 is completely absent among early agriculturalist DNA samples from Neolithic Scandinavia.[41][42][32] Except for a single DNA sample (SF11), it is also absent from Mesolithic hunter-gatherers in Scandinavia. I1 first starts to appear in Scandinavia in notable frequency during the late Neolithic in conjunction with the entrance of groups carrying Western Steppe Herder ancestry into Scandinavia, but does not increase significantly in frequency until the beginning of the Nordic Bronze Age.[37][43][44]
Due to the very low number of ancient DNA samples that have been assigned to I1 that date to earlier than the Nordic Bronze Age, it is currently unknown whether I1 was present as a rare haplogroup among Scandinavian forager cultures such as Pitted Ware before becoming assimilated by the Battle Axe culture, or if it was brought into Scandinavia by incoming groups such as Battle Axe who may have assimilated it from cultures such as the Funnelbeaker culture in Central Europe; or the steppe itself. Future research will most likely be able to determine which one of these two possible origins turns out to be the case.
Samples SF11 and BAB5 are unlike other ancient DNA samples assigned to I1 in the sense that they both seem to represent now-extinct branches of I1 that hadn't fully developed into I-M253 yet. They are therefore unlikely to have been ancestral to present-day carriers of I1, who all share a common ancestor that lived around 2600 BC.
According to a study published in 2010, I-M253 originated between 3,170 and 5,000 years ago, in Chalcolithic Europe.[2] A new study in 2015 estimated the origin as between 3,470 and 5,070 years ago or between 3,180 and 3,760 years ago, using two different techniques.[3]
In 2007, it was suggested that it initially dispersed from the area that is now Denmark.[14] However, Prof. Dr. Kenneth Nordtvedt, Montana State University, regarding the MRCA, in 2009 wrote in a personal message: "We don't know where that man existed, but the greater lower Elbe basin seems like the heartland of I1".
Latest results (January 2022) published by Y-Full suggest I1 (I-M253) was formed 27,500 ybp (95 CI: 29,800 ybp – 25,200 ybp) with TMRCA 4,600 ybp (95 CI: 5,200 ybp – 4,000 ybp). Since the most up-to date calculated estimation of TMRCA of I1 is thought to be around 2600 BC, this likely puts the ancestor of all living I1 men somewhere in Northern Europe around that time. The phylogeny of I1 shows the signature of a rapid star-like expansion.[45][46] This suggests that I1 went from being a rare marker to a rather common one in a rapid burst.[3]
Structure
[edit]I-M253 (M253, M307.2/P203.2, M450/S109, P30, P40, L64, L75, L80, L81, L118, L121/S62, L123, L124/S64, L125/S65, L157.1, L186, and L187) or I1 [7]
- I-DF29 (DF29/S438); I1a
- I-CTS6364 (CTS6364/Z2336); I1a1
- FGC20030; I1a1a~
- S4767; I1a1a1~
- I-M227; I1a1a1a1a
- A394; I1a1a2~
- Y11221; I1a1a3~
- A5338; I1a1a4~
- S4767; I1a1a1~
- CTS10028; I1a1b
- I-L22 (L22/S142); I1a1b1
- CTS11651/Z2338; I1a1b1a~
- I-P109; I1a1b1a1
- I-Y3662; I1a1b1a1e~
- I-S14887; I1a1b1a1e2~
- I-Y11203; I1a1b1a1e2d~
- I-Y29630; I1a1b1a1e2d2~
- I-Y11203; I1a1b1a1e2d~
- I-S14887; I1a1b1a1e2~
- I-Y3662; I1a1b1a1e~
- CTS6017; I1a1b1a2
- I-L205 (L205.1/L939.1/S239.1); I1a1b1a3
- CTS6868; I1a1b1a4
- I-Z74; I1a1b1a4a
- CTS2208; I1a1b1a4a1~
- I-L287; I1a1b1a4a1a
- I-L258 (L258/S335); I1a1b1a4a1a1
- I-L287; I1a1b1a4a1a
- I-L813; I1a1b1a4a2
- FGC12562; I1a1b1a4a3~
- CTS2208; I1a1b1a4a1~
- I-Z74; I1a1b1a4a
- I-P109; I1a1b1a1
- CTS11603/S4744; I1a1b1b~
- I-FT40464
- I-Y19934
- I-L300 (L300/S241); I1a1b1b1a1
- I-Y31032
- I-Y32014
- I-Y22918
- I-Y21972
- I-Y24013
- I-Y24015
- I-Y31032
- I-Y19933
- I-Y19932
- I-Y22015
- I-FT57000
- I-Y22015
- I-Y19932
- I-L300 (L300/S241); I1a1b1b1a1
- I-Y19934
- I-FT40464
- CTS11651/Z2338; I1a1b1a~
- FGC10477/Y13056; I1a1b2
- A8178, A8182, A8200, A8204; I1a1b3~
- F13534.2/Y17263.2; I1a1b4~
- I-L22 (L22/S142); I1a1b1
- FGC20030; I1a1a~
- I-Z58 (S244/Z58); I1a2
- I-Z59 (S246/Z59); I1a2a
- I-Z60 (S337/Z60, S439/Z61, Z62); I1a2a1
- I-Z140 (Z140, Z141)
- I-L338
- I-F2642 (F2642)
- I-Z73
- I-L1302
- I-L573
- I-L803
- I-Z140 (Z140, Z141)
- I-Z382; I1a2a2
- I-Z60 (S337/Z60, S439/Z61, Z62); I1a2a1
- I-Z138 (S296/Z138, Z139); I1a2b
- I-Z2541
- I-Z59 (S246/Z59); I1a2a
- I-Z63 (S243/Z63); I1a3
- I-BY151; I1a3a
- I-L849.2; I1a3a1
- I-BY351; I1a3a2
- I-CTS10345
- I-Y10994
- I-Y7075
- I-CTS10345
- I-S2078
- I-S2077
- I-Y2245 (Y2245/PR683)
- I-L1237
- I-FGC9550
- I-S10360
- I-S15301
- I-Y7234
- I-L1237
- I-Y2245 (Y2245/PR683)
- I-S2077
- I-BY62 (BY62); I1a3a3
- I-BY151; I1a3a
- I-CTS6364 (CTS6364/Z2336); I1a1
- I-Z131 (Z131/S249); I1b
- I-CTS6397; I1b1
- I-Z17943 (Y18119/Z17925, S2304/Z17937); I1c
Historical expansion
[edit]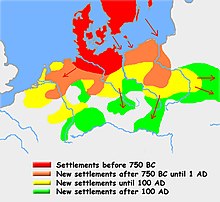
Haplogroup I1, as well as subclades of R1b such as R1b-Z18 and subclades of R1a such as R1a-Z284, are strongly associated with Germanic peoples and are linked to the proto-Germanic speakers of the Nordic Bronze Age.[47][48] Current DNA research indicates that I1 was close to non-existent in most of Europe outside of Scandinavia and northern Germany before the Migration Period. The expansion of I1 is directly tied to that of the Germanic tribes. Starting around 900 BC, Germanic tribes started moving out of southern Scandinavia and northern Germany into the nearby lands between the Elbe and the Oder. Between 600 and 300 BC another wave of Germanic peoples migrated across the Baltic Sea and settled alongside the Vistula. Germanic migration to that area resulted in the formation of the Wielbark culture, which is associated with the Goths.[49]
I1-Z63 has been traced to the Kowalewko burial site in Poland which dates to the Roman Iron Age. In 2017 Polish researchers could successfully assign YDNA haplogroups to 16 individuals who were buried at the site. Out of these 16 individuals, 8 belonged to I1. In terms of subclades, three belonged to I-Z63, and in particular subclade I-L1237.[50] The Kowalewko archeological site has been associated with the Wielbark culture. Therefore, the subclade I-L1237 of I-Z63 may be seen somewhat as a genetic indicator of the Gothic tribe of late antiquity. I1-Z63 has also been found in a burial associated with Goth and Lombard remains in Collegno, Italy.[51][52] The cemetery is dated to the late 6th Century and further suggests that I1-Z63 and downstream subclades are linked to early Medieval Gothic migrations.
In 2015, a DNA study tested the Y-DNA haplogroups of 12 samples dated to 300–400 AD from Saxony-Anhalt in Germany. 8 of them belonged to haplogroup I1. This DNA evidence is in alignment with the historical migrations of Germanic tribes from Scandinavia to central Europe.[53]
Additionally, I1-Z63 was found in the Late Antiquity site Crypta Balbi in Rome, this time with the downstream subclade I-Y7234.[54] Material findings associated with the Lombards have been excavated in Crypta Balbi.
The Pla de l'Horta villa near Girona in Spain is located in close proximity to a necropolis with a series of tombs associated with the Visigoths. The grave goods and the typology of the tombs point to a Visigothic origin of the individuals. A small number of individuals buried at the site were sampled for DNA analysis in a 2019 study. One of the samples belonged to haplogroup I1.[55] This finding is in accordance with the common ancestral origin of the Visigoths and the Ostrogoths.
The Anglo-Saxon settlement of Britain introduced I1 in the British Isles.[56] A 2022 study found that out of 120 samples from Anglo-Saxon period England, 41 samples or roughly 34.17% of the samples belonged to haplogroup I1. The study noted that there was a heavy correlation between "CNE" Continental North European-like ancestry and Y-DNA I1.[57]
During the Viking Age, I-M253 saw another expansion. Margaryan et al. 2020 analyzed 442 Viking world individuals from various archaeological sites in Europe. I-M253 was the most common Y-haplogroup found in the study. Norwegian and Danish Vikings brought more I1 to Britain and Ireland, while Swedish Vikings introduced it to Russia and Ukraine and brought more of it to Finland and Estonia.[58]
Geographical distribution
[edit]I-M253 is found at its highest density in Northern Europe and other countries that experienced extensive migration from Northern Europe, either in the Migration Period, the Viking Age, or modern times. It is found in all places invaded by the Norse.
During the modern era, significant I-M253 populations have also taken root in immigrant nations and former European colonies such as the United States, Australia, New Zealand and Canada.
| Population | Sample size | I (total) | I1 (I-M253) | I1a1a (I-M227) | Source |
|---|---|---|---|---|---|
| Albanians (Tirana) | 55 | 21.82%=(12/55) | 3.6%=(2/55) | 0.0 | Battaglia et al. 2008 |
| Albanians (North Macedonia) | 64 | 17.2%=(11/64) | 4.7%=(3/64) | 0.0 | Battaglia et al. 2008 |
| Albanians (Tirana) Albanians (North Macedonia) |
55+64=119 | 19.33%=(23/119) | 4.2%=(5/119) | 0.0 | Battaglia et al. 2008 |
| Kosovo Albanians (Pristina) | 114 | 7.96%=(9/114) | 5.31%=(6/114) | 0.0 | Pericic et al. 2005 |
| Albanians (Tirana) Albanians (North Macedonia) Kosovo Albanians (Pristina) |
55+64+114=233 | 13.73%=(32/233) | 4.72%=(11/233) | 0.0 | Pericic et al. 2005 Battaglia et al. 2008 |
| Austria | 43 | 9.3 | 2.3 | 0.0 | Underhill et al. 2007 |
| Belarus: Vitbsk | 100 | 15 | 1.0 | 0.0 | Underhill et al. 2007 |
| Belarus: Brest | 97 | 20.6 | 1.0 | 0.0 | Underhill et al. 2007 |
| Bosnia | 100 | 42 | 2.0 | 0.0 | Rootsi et al. 2004 |
| Bulgaria | 808 | 26.6 | 4.3 | 0.0 | Karachanak et al. 2013 |
| Czech Republic | 47 | 31.9 | 8.5 | 0.0 | Underhill et al. 2007 |
| Czech Republic | 53 | 17.0 | 1.9 | 0.0 | Rootsi et al. 2004 |
| Denmark | 122 | 39.3% (48/122) | 34.8% (40/122) | 0.0 | Underhill et al. 2007 |
| England | 104 | 19.2 | 15.4 | 0.0 | Underhill et al. 2007 |
| Estonia | 210 | 18.6 | 14.8 | 0.5 | Rootsi et al. 2004 |
| Estonia | 118 | 11.9 | Lappalainen et al. 2008 | ||
| Finland (national) | 28.0 | Lappalainen et al. 2006 | |||
| Finland: West | 230 | 40% (92/230) | Lappalainen et al. 2008 | ||
| Finland: East | 306 | 19% (58/306) | Lappalainen et al. 2008 | ||
| Finland: Satakunta region | 50+ | Lappalainen et al. 20089 | |||
| France | 58 | 17.2 | 8.6 | 1.7 | Underhill et al. 2007 |
| France | 12 | 16.7 | 16.7 | 0.0 | Cann et al. 2002 |
| France (Low Normandy) | 42 | 21.4 | 11.9 | 0.0 | Rootsi et al. 2004 |
| Germany | 125 | 24 | 15.2 | 0.0 | Underhill et al. 2007 |
| Greece | 171 | 15.8 | 2.3 | 0.0 | Underhill et al. 2007 |
| Hungary | 113 | 25.7 | 13.3 | 0.0 | Rootsi et al. 2004 |
| Ireland | 100 | 11 | 6.0 | 0.0 | Underhill et al. 2007 |
| Volga Tatars | 53 | 13.2 | 11.3 | 0.0 | Trofimova 2015 |
| Latvia | 113 | 3.5 | Lappalainen et al. 2008 | ||
| Lithuania | 164 | 4.9 | Lappalainen et al. 2008 | ||
| Netherlands | 93 | 20.4 | 14 | 0.0 | Underhill et al. 2007 |
| Norway | 1766 | 37% (653/1766) | Stenersen et al. 2006 | ||
| Russia (national) | 16 | 25 | 12.5 | 0.0 | Cann et al. 2002 |
| Russia: Pskov | 130 | 16.9 | 5.4 | 0.0 | Underhill et al. 2007 |
| Russia: Kostroma | 53 | 26.4 | 11.3 | 0.0 | Underhill et al. 2007 |
| Russia: Smolensk | 103 | 12.6 | 1.9 | 0.0 | Underhill et al. 2007 |
| Russia: Voronez | 96 | 19.8 | 3.1 | 0.0 | Underhill et al. 2007 |
| Russia: Arkhangelsk | 145 | 15.8 | 7.6 | 0.0 | Underhill et al. 2007 |
| Russia: Cossacks | 89 | 24.7 | 4.5 | 0.0 | Underhill et al. 2007 |
| Russia: Karelians | 140 | 10 | 8.6 | 0.0 | Underhill et al. 2007 |
| Russia: Karelians | 132 | 15.2 | Lappalainen et al. 2008 | ||
| Russia: Vepsa | 39 | 5.1 | 2.6 | 0.0 | Underhill et al. 2007 |
| Slovakia | 70 | 14.3 | 4.3 | 0.0 | Rootsi et al. 2004 |
| Slovenia | 95 | 26.3 | 7.4 | 0.0 | Underhill et al. 2007 |
| Sweden (national) | 160 | 35.6% (57/160) | Lappalainen et al. 2008 | ||
| Sweden (national) | 38.0 | Lappalainen et al. 2009 | |||
| Sweden: Västra Götaland | 52 | Lappalainen et al. 2009 | |||
| Switzerland | 144 | 7.6 | 5.6 | 0.0 | Rootsi et al. 2004 |
| Turkey | 523 | 5.4 | 1.1 | 0.0 | Underhill et al. 2007 |
| Ukraine: Lviv | 101 | 23.8 | 4.9 | 0.0 | Underhill et al. 2007 |
| Ukraine: Ivanovo-Frankiv | 56 | 21.4 | 1.8 | 0.0 | Underhill et al. 2007 |
| Ukraine: Hmelnitz | 176 | 26.2 | 6.1 | 0.0 | Underhill et al. 2007 |
| Ukraine: Cherkasy | 114 | 28.1 | 4.3 | 0.0 | Underhill et al. 2007 |
| Ukraine: Bilhorod | 56 | 26.8 | 5.3 | 0.0 | Underhill et al. 2007 |
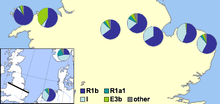
In 2002 a paper was published by Michael E. Weale and colleagues showing genetic evidence for population differences between the English and Welsh populations, including a markedly higher level of Y-DNA haplogroup I1 in England than in Wales. They saw this as convincing evidence of Anglo-Saxon mass invasion of eastern Great Britain from northern Germany and Denmark during the Migration Period.[59] The authors assumed that populations with large proportions of haplogroup I1 originated from northern Germany or southern Scandinavia, particularly Denmark, and that their ancestors had migrated across the North Sea with Anglo-Saxon migrations and Danish Vikings. The main claim by the researchers was
that an Anglo-Saxon immigration event affecting 50–100% of the Central English male gene pool at that time is required. We note, however, that our data do not allow us to distinguish an event that simply added to the indigenous Central English male gene pool from one where indigenous males were displaced elsewhere or one where indigenous males were reduced in number ... This study shows that the Welsh border was more of a genetic barrier to Anglo-Saxon Y chromosome gene flow than the North Sea ... These results indicate that a political boundary can be more important than a geophysical one in population genetic structuring.
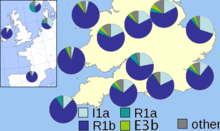
In 2003 a paper was published by Christian Capelli and colleagues which supported, but modified, the conclusions of Weale and colleagues.[60] This paper, which sampled Great Britain and Ireland on a grid, found a smaller difference between Welsh and English samples, with a gradual decrease in Haplogroup I1 frequency moving westwards in southern Great Britain. The results suggested to the authors that Norwegian Vikings invaders had heavily influenced the northern area of the British Isles, but that both English and mainland Scottish samples all have German/Danish influence.
Prominent members of I-M253
[edit]Through direct testing or testing of their descendants and genealogical evidence, the following notable people have been shown to be I-M253:
- Alexander Hamilton, founding father of the United States.[61]
- The Varangian Šimon, said to be the founder of the Russian Vorontsov noble family (including Prince Mikhail Semyonovich Vorontsov and Illarion Ivanovich Vorontsov-Dashkov) belonged to haplogroup I1-Y15024.[62][63][64][65]
- The Rurikid Prince Sviatopolk the Accursed (son of Vladimir the Great) belonged to I1-S2077.[66][67][68]
- Birger Jarl, Duke of Sweden of the East Geatish House of Bjälbo, founder of Stockholm; his remains were exhumed and tested in 2002 and found to be I-M253.[69] The House of Bjälbo also provided three kings of Norway, and one king of Denmark in the 14th century.
- British musician Gordon Sumner, known as Sting[70]
- William Bradford, Governor of the Plymouth Colony[71]
- William Brewster, an early colonist who emigrated to America on the Mayflower[71]
- Confederate general Robert E. Lee. Other prominent members of the Lee family of Virginia and Maryland were Richard Lee I and Richard Henry Lee.[72]
- The House of Grimaldi belong to a Scandinavian subclade of I1, downstream of I1-Y3549.[73]
- President of the United States Andrew Jackson.[74][75]
- Russian writer Leo Tolstoy.[76][77]
- Icelandic historian and poet Snorri Sturluson[76]
- Swedish scientist and theologian Emanuel Swedenborg[78][79]
- Siener van Rensburg, Boer patriotic figure and mystic.[80][81]
- Björn Wahlroos, Finnish businessman and millionaire.[76]
- Finnish mathematician Rolf Nevanlinna.[82][83][84]
- American inventor Samuel Morse.[85][86][87]
- Swedish Footballers Sebastian Larsson and his father Svante Larsson belong to I1-Y24470.[88][89][90][91]
- Swedish YouTuber Felix Kjellberg (PewDiePie) belongs to I1-L22.[92]
- Swedish actor Björn Andrésen belongs to haplogroup I1-L22.[93][94][95][96] His ancestor Johan Andrésen lived on both sides of the Swedish-Norwegian border.[97][98]
- American actor Chris Pine belongs to haplogroup I1-A13819.[99][100]
- Swedish Sámi Ice hockey player Börje Salming.[101]
- American colonist Edmund Rice.
- German composer Ludwig van Beethoven.[102]
Markers
[edit]
The following are the technical specifications for known I-M253 haplogroup SNP and STR mutations.
Name: M253[103]
- Type: SNP
- Source: M (Peter Underhill of Stanford University)
- Position: ChrY:13532101..13532101 (+ strand)
- Position (base pair): 283
- Total size (base pairs): 400
- Length: 1
- ISOGG HG: I1
- Primer F (Forward 5′→ 3′): GCAACAATGAGGGTTTTTTTG
- Primer R (Reverse 5′→ 3′): CAGCTCCACCTCTATGCAGTTT
- YCC HG: I1
- Nucleotide alleles change (mutation): C to T
Name: M307[104]
- Type: SNP
- Source: M (Peter Underhill)
- Position: ChrY:21160339..21160339 (+ strand)
- Length: 1
- ISOGG HG: I1
- Primer F: TTATTGGCATTTCAGGAAGTG
- Primer R: GGGTGAGGCAGGAAAATAGC
- YCC HG: I1
- Nucleotide alleles change (mutation): G to A
Name: P30[105]
- Type: SNP
- Source: PS (Michael Hammer of the University of Arizona and James F. Wilson, at the University of Edinburgh)
- Position: ChrY:13006761..13006761 (+ strand)
- Length: 1
- ISOGG HG: I1
- Primer F: GGTGGGCTGTTTGAAAAAGA
- Primer R: AGCCAAATACCAGTCGTCAC
- YCC HG: I1
- Nucleotide alleles change (mutation): G to A
- Region: ARSDP
Name: P40[106]
- Type: SNP
- Source: PS (Michael Hammer and James F. Wilson)
- Position: ChrY:12994402..12994402 (+ strand)
- Length: 1
- ISOGG HG: I1
- Primer F: GGAGAAAAGGTGAGAAACC
- Primer R: GGACAAGGGGCAGATT
- YCC HG: I1
- Nucleotide alleles change (mutation): C to T
- Region: ARSDP
See also
[edit]References
[edit]- ^ a b c "I1 YTree". Yfull.com. 2022-04-06. Retrieved 2022-04-19.
- ^ a b Pedro Soares, Alessandro Achilli, Ornella Semino, William Davies, Vincent Macaulay, Hans-Jürgen Bandelt, Antonio Torroni, and Martin B. Richards, The Archaeogenetics of Europe, Current Biology, vol. 20 (February 23, 2010), R174–R183. yDNA Haplogroup I: Subclade I1, Family Tree DNA,
- ^ a b c Batini C, Hallast P, Zadik D, Delser PM, Benazzo A, Ghirotto S, et al. (May 2015). "Large-scale recent expansion of European patrilineages shown by population resequencing". Nature Communications. 6: 7152. Bibcode:2015NatCo...6.7152B. doi:10.1038/ncomms8152. PMC 4441248. PMID 25988751.
- ^ Rootsi S, Magri C, Kivisild T, Benuzzi G, Help H, Bermisheva M, et al. (July 2004). "Phylogeography of Y-chromosome haplogroup I reveals distinct domains of prehistoric gene flow in europe" (PDF). American Journal of Human Genetics. 75 (1): 128–37. doi:10.1086/422196. PMC 1181996. PMID 15162323. Archived from the original (PDF) on 2009-06-24. Retrieved 2008-03-20.
- ^ Underhill PA, Myres NM, Rootsi S, Chow CT, Lin AA, Otillar RP, et al. (2007). "New phylogenetic relationships for Y-chromosome haplogroup I: reappraising its phylogeography and prehistory.". In Mellars P, Boyle K, Bar-Yosef O, Stringe C (eds.). Rethinking the Human Revolution. Cambridge, UK: McDonald Institute Monographs. pp. 33–42. ISBN 978-1-902937-46-5.
- ^ Lappalainen, T.; Laitinen, V.; Salmela, E.; Andersen, P.; Huoponen, K.; Savontaus, M.-L.; Lahermo, P. (2008). "Migration Waves to the Baltic Sea Region". Annals of Human Genetics. 72 (3): 337–348. doi:10.1111/j.1469-1809.2007.00429.x. PMID 18294359. S2CID 32079904.
- ^ a b ISOGG, Y-DNA Haplogroup I and its Subclades – 2017 (31 January 2017).
- ^ a b Lappalainen T, Laitinen V, Salmela E, Andersen P, Huoponen K, Savontaus ML, Lahermo P (May 2008). "Migration waves to the Baltic Sea region". Annals of Human Genetics. 72 (Pt 3): 337–348. doi:10.1111/j.1469-1809.2007.00429.x. PMID 18294359. S2CID 32079904.
- ^ Lappalainen T, Hannelius U, Salmela E, von Döbeln U, Lindgren CM, Huoponen K, et al. (January 2009). "Population structure in contemporary Sweden—a Y-chromosomal and mitochondrial DNA analysis". Annals of Human Genetics. 73 (1): 61–73. doi:10.1111/j.1469-1809.2008.00487.x. PMID 19040656. S2CID 205598345.
- ^ "FamilyTreeDNA – Sweden DNA PROJECT – Sverigeprojektet".
- ^ Dupuy BM, Stenersen M, Lu TT, Olaisen B (December 2006). "Geographical heterogeneity of Y-chromosomal lineages in Norway". Forensic Science International. 164 (1): 10–19. doi:10.1016/j.forsciint.2005.11.009. PMID 16337760.
- ^ "FamilyTreeDNA – The Norway DNA Project – Norgesprosjektet". familytreedna.com. Retrieved 2020-11-26.
- ^ "Y-DNA Haplogrupper". Norway DNA Norgesprosjektet. Retrieved 2020-12-27.
- ^ a b Peter A. Underhill et al., New Phylogenetic Relationships for Y-chromosome Haplogroup I: Reappraising its Phylogeography and Prehistory, in Rethinking the Human Revolution (2007), pp. 33–42. P. Mellars, K. Boyle, O. Bar-Yosef, C. Stringer (Eds.) McDonald Institute for Archaeological Research, Cambridge, UK.
- ^ Sanchez JJ (2004). "Y chromosome SNP haplogroups in Danes, Greenlanders and Somalis" (PDF). International Congress Series. 1261: 347–49. doi:10.1016/S0531-5131(03)01635-2 – via isfg.org.
- ^ "FamilyTreeDNA – Denmark DNA Project". familytreedna.com. Retrieved 2020-12-10.
- ^ Helgason A, Sigureth ardóttir S, Nicholson J, Sykes B, Hill EW, Bradley DG, et al. (September 2000). "Estimating Scandinavian and Gaelic ancestry in the male settlers of Iceland". American Journal of Human Genetics. 67 (3): 697–717. doi:10.1086/303046. PMC 1287529. PMID 10931763.
- ^ "The Greater Nordic Regional Y-DNA Project". familytreedna. April 2021.
- ^ Ebenesersdóttir SS, Sandoval-Velasco M, Gunnarsdóttir ED, Jagadeesan A, Guðmundsdóttir VB, Thordardóttir EL, et al. (June 2018). "Ancient genomes from Iceland reveal the making of a human population". Science. 360 (6392): 1028–1032. Bibcode:2018Sci...360.1028E. doi:10.1126/science.aar2625. hdl:10852/71890. PMID 29853688.
- ^ Lappalainen T, Koivumäki S, Salmela E, Huoponen K, Sistonen P, Savontaus ML, Lahermo P (July 2006). "Regional differences among the Finns: a Y-chromosomal perspective". Gene. 376 (2): 207–15. doi:10.1016/j.gene.2006.03.004. PMID 16644145.
- ^ "Blood of the Isles: exploring the genetic roots of our tribal history". History Ireland. 2013-02-22. Retrieved 2020-12-10.
- ^ Karafet TM, Mendez FL, Meilerman MB, Underhill PA, Zegura SL, Hammer MF (May 2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. 18 (5): 830–38. doi:10.1101/gr.7172008. PMC 2336805. PMID 18385274.
- ^ Batini C, Hallast P, Zadik D, Delser PM, Benazzo A, Ghirotto S, et al. (May 2015). "Large-scale recent expansion of European patrilineages shown by population resequencing". Nature Communications. 6 (1): 7152. Bibcode:2015NatCo...6.7152B. doi:10.1038/ncomms8152. PMC 4441248. PMID 25988751.
- ^ a b "Welcome to FamilyTreeDNA Discover (Beta)". FamilyTreeDNA Discover (Beta). Retrieved 2022-12-25.
- ^ "I-DF29 YTree". yfull.com. Retrieved 2022-12-25.
- ^ Malmström H, Günther T, Svensson EM, Juras A, Fraser M, Munters AR, et al. (October 2019). "The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon". Proceedings. Biological Sciences. 286 (1912): 20191528. doi:10.1098/rspb.2019.1528. PMC 6790770. PMID 31594508.
- ^ "I-Y11204 YTree". yfull.com. Retrieved 2022-12-25.
- ^ Allentoft, Morten E.; Sikora, Martin; Fischer, Anders; Sjögren, Karl-Göran; Ingason, Andrés; Macleod, Ruairidh; Rosengren, Anders; Schulz Paulsson, Bettina; Jørkov, Marie Louise Schjellerup; Novosolov, Maria; Stenderup, Jesper; Price, T. Douglas; Fischer Mortensen, Morten; Nielsen, Anne Birgitte; Ulfeldt Hede, Mikkel (January 2024). "100 ancient genomes show repeated population turnovers in Neolithic Denmark". Nature. 625 (7994): 329–337. doi:10.1038/s41586-023-06862-3. ISSN 1476-4687. PMC 10781617.
- ^ "Haplotree.info – ancientdna.info. Map based on All Ancient DNA v. 2.07.26".
- ^ Gunther T (2017). "Genomics of Mesolithic Scandinavia reveal colonization routes and high-latitude adaptation" (PDF). Nature. 23: 6 – via Biorxiv.
- ^ SF11 – Stora Förvar, Stora Karlsö I-Z2699*. "Haplotree.info sample: SF11". haplotree.info.
{{cite web}}: CS1 maint: numeric names: authors list (link) - ^ a b Skoglund P, Malmström H, Omrak A, Raghavan M, Valdiosera C, Günther T, et al. (May 2014). "Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers". Science. 344 (6185): 747–50. Bibcode:2014Sci...344..747S. doi:10.1126/science.1253448. PMID 24762536. S2CID 206556994.
- ^ "familytreedna.com I-Z2699 tree". familytreedna. April 2021.
- ^ Szécsényi-Nagy A, Brandt G, Haak W, Keerl V, Jakucs J, Möller-Rieker S, et al. (April 2015). "Tracing the genetic origin of Europe's first farmers reveals insights into their social organization". Proceedings. Biological Sciences. 282 (1805). doi:10.1098/rspb.2015.0339. PMC 4389623. PMID 25808890.
- ^ "BAB5 I-Z2699*". haplotree.info.
- ^ Vyas, Deven N.; Koncz, István; Modi, Alessandra; Mende, Balázs Gusztáv; Tian, Yijie; Francalacci, Paolo; Lari, Martina; Vai, Stefania; Straub, Péter; Gallina, Zsolt; Szeniczey, Tamás; Hajdu, Tamás; Pejrani Baricco, Luisella; Giostra, Caterina; Radzevičiūtė, Rita (2023-09-25). "Fine-scale sampling uncovers the complexity of migrations in 5th-6th century Pannonia". Current biology: CB. 33 (18): 3951–3961.e11. doi:10.1016/j.cub.2023.07.063. hdl:11584/373523. ISSN 1879-0445. PMID 37633281.
- ^ a b c Allentoft ME, Sikora M, Sjögren KG, Rasmussen S, Rasmussen M, Stenderup J, et al. (June 2015). "Population genomics of Bronze Age Eurasia". Nature. 522 (7555): 167–72. Bibcode:2015Natur.522..167A. doi:10.1038/nature14507. PMID 26062507. S2CID 4399103.
- ^ "YFull | The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon". yfull.com. Retrieved 2021-01-24.
- ^ Malmstrom H (July 2019). "The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon" (PDF). Uppsala Genomics. 1: 3 – via jakobssonlab.iob.uu.se/.
- ^ "I-Y11204 YTree".
- ^ Sánchez-Quinto F, Malmström H, Fraser M, Girdland-Flink L, Svensson EM, Simões LG, et al. (May 2019). "Megalithic tombs in western and northern Neolithic Europe were linked to a kindred society". Proceedings of the National Academy of Sciences of the United States of America. 116 (19): 9469–74. Bibcode:2019PNAS..116.9469S. doi:10.1073/pnas.1818037116. PMC 6511028. PMID 30988179.
- ^ Malmström H, Linderholm A, Skoglund P, Storå J, Sjödin P, Gilbert MT, et al. (January 2015). "Ancient mitochondrial DNA from the northern fringe of the Neolithic farming expansion in Europe sheds light on the dispersion process". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 370 (1660): 20130373. doi:10.1098/rstb.2013.0373. PMC 4275881. PMID 25487325.
- ^ Karlsson AO, Wallerström T, Götherström A, Holmlund G (August 2006). "Y-chromosome diversity in Sweden – a long-time perspective". European Journal of Human Genetics. 14 (8): 963–70. doi:10.1038/sj.ejhg.5201651. PMID 16724001. S2CID 23227271.
- ^ Malmström H, Günther T, Svensson EM, Juras A, Fraser M, Munters AR, et al. (October 2019). "The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon". Proceedings. Biological Sciences. 286 (1912): 20191528. doi:10.1098/rspb.2019.1528. PMC 6790770. PMID 31594508.
- ^ Poznik GD, Xue Y, Mendez FL, Willems TF, Massaia A, Wilson Sayres MA, et al. (June 2016). "Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences". Nature Genetics. 48 (6): 593–99. doi:10.1038/ng.3559. hdl:11858/00-001M-0000-002A-F024-C. PMC 4884158. PMID 27111036.
- ^ Woodley M (February 2017). "Holocene selection for variants associated with cognitive ability: Comparing ancient and modern genomes" (PDF). biorxiv.org/. doi:10.1101/109678. S2CID 196631764. Retrieved 27 January 2021.
- ^ Schmidt KH (1991). "The Celts and the Ethnogenesis of the Germanic People". Historische Sprachforschung / Historical Linguistics. 104 (1): 129–52. ISSN 0935-3518. JSTOR 40849016.
- ^ Bergerbrant S (May 2007). "Bronze Age Identities: Costume, Conflict and Contact in Northern Europe 1600–1300 BC" (PDF). Stockholm Studies in Archaeology. 43: 7–201 – via diva-portal.org.
- ^ Teska M, Michalowski A (2014). "Connection between Wielkopolska and the Baltic Sea Region in the Roman Iron". Archaeologia Lituana. 14: 63–77. doi:10.15388/ArchLit.2013.0.2641. S2CID 56295624.
- ^ Zenczak M, Handschuh L, Juras A, Marcinkowska-Swojak M, Philips A, Piontek J, Stolarek I, Figlerowicz M (2017). Y-Chromosome Haplogroup Assignment Through Next Generation Sequencing of Enriched Ancient DNA Libraries. Anthropological Genetics. p. Presentation number: AG 16.
- ^ Amorim CE, Vai S, Posth C, Modi A, Koncz I, Hakenbeck S, et al. (September 2018). "Understanding 6th-century barbarian social organization and migration through paleogenomics". Nature Communications. 9 (1): 3547. Bibcode:2018NatCo...9.3547A. doi:10.1038/s41467-018-06024-4. PMC 6134036. PMID 30206220.
- ^ Estes R (2020-10-16). "Longobards Ancient DNA from Pannonia and Italy – What Does Their DNA Tell Us? Are You Related?". DNAeXplained – Genetic Genealogy. Retrieved 2020-12-11.
- ^ Labudde D (July 2015). "Gender distribution of excavation finds from the Roman imperial and migration period". ResearchGate. 1: 2 – via ResearchGate.net.
- ^ Antonio ML, Gao Z, Moots HM, Lucci M, Candilio F, Sawyer S, et al. (November 2019). "Ancient Rome: A genetic crossroads of Europe and the Mediterranean". Science. 366 (6466): 708–14. Bibcode:2019Sci...366..708A. doi:10.1126/science.aay6826. PMC 7093155. PMID 31699931.
- ^ Olalde I, Mallick S, Patterson N, Rohland N, Villalba-Mouco V, Silva M, et al. (March 2019). "The genomic history of the Iberian Peninsula over the past 8000 years". Science. 363 (6432): 1230–34. Bibcode:2019Sci...363.1230O. doi:10.1126/science.aav4040. PMC 6436108. PMID 30872528.
- ^ Martiniano R, Caffell A, Holst M, Hunter-Mann K, Montgomery J, Müldner G, et al. (January 2016). "Genomic signals of migration and continuity in Britain before the Anglo-Saxons". Nature Communications. 7 (1): 10326. Bibcode:2016NatCo...710326M. doi:10.1038/ncomms10326. PMC 4735653. PMID 26783717.
- ^ Gretzinger, Joscha; Sayer, Duncan; Justeau, Pierre; Altena, Eveline; Pala, Maria; Dulias, Katharina; Edwards, Ceiridwen J.; Jodoin, Susanne; Lacher, Laura; Sabin, Susanna; Vågene, Åshild J.; Haak, Wolfgang; Ebenesersdóttir, S. Sunna; Moore, Kristjan H. S.; Radzeviciute, Rita (October 2022). "The Anglo-Saxon migration and the formation of the early English gene pool". Nature. 610 (7930): 112–119. doi:10.1038/s41586-022-05247-2. ISSN 1476-4687. PMC 9534755.
- ^ Margaryan A, Lawson DJ, Sikora M, Racimo F, Rasmussen S, Moltke I, et al. (September 2020). "Population genomics of the Viking world". Nature. 585 (7825): 390–96. Bibcode:2020Natur.585..390M. bioRxiv 10.1101/703405. doi:10.1038/s41586-020-2688-8. hdl:10852/83989. PMID 32939067. S2CID 201195157.
- ^ Weale ME, Weiss DA, Jager RF, Bradman N, Thomas MG (July 2002). "Y chromosome evidence for Anglo-Saxon mass migration". Molecular Biology and Evolution. 19 (7): 1008–21. doi:10.1093/oxfordjournals.molbev.a004160. PMID 12082121.
- ^ Capelli C, Redhead N, Abernethy JK, Gratrix F, Wilson JF, Moen T, et al. (May 2003). "A Y chromosome census of the British Isles". Current Biology. 13 (11): 979–84. doi:10.1016/S0960-9822(03)00373-7. hdl:20.500.11820/8acb01f3-a7c1-45f5-89de-b796266d651e. PMID 12781138. S2CID 526263.
- ^ "Founding Father DNA". isogg.org.
- ^ "FamilyTreeDNA – Genetic Testing for Ancestry, Family History & Genealogy". familytreedna.com. Retrieved 2020-12-10.
- ^ "Faderslinjen, DNA". sikaby.se. Retrieved 2020-12-10.
- ^ "FamilyTreeDNA – RussiaDNA Project". familytreedna.com. Retrieved 2020-12-10.
- ^ "Vår vikingahövding i österled". sikaby.se. Retrieved 2020-12-10.
- ^ "Sample from Homo sapiens – BioSample – NCBI". ncbi.nlm.nih.gov. Retrieved 2020-12-10.
- ^ Margaryan A, Lawson DJ, Sikora M, Racimo F, Rasmussen S, Moltke I, et al. (September 2020). "Population genomics of the Viking world". Nature. 585 (7825): 390–96. Bibcode:2020Natur.585..390M. doi:10.1038/s41586-020-2688-8. hdl:10852/83989. PMID 32939067. S2CID 221769227.
- ^ Duczko W (2004). Viking Rus: Studies on the Presence of Scandinavians in Eastern Europe. Brill. ISBN 978-90-04-13874-2.
- ^ Malmström H, Vretemark M, Tillmar A, Durling MB, Skoglund P, Gilbert MT, et al. (January 2012). "Finding the founder of Stockholm – a kinship study based on Y-chromosomal, autosomal and mitochondrial DNA". Annals of Anatomy - Anatomischer Anzeiger. Special Issue: Ancient DNA. 194 (1): 138–45. doi:10.1016/j.aanat.2011.03.014. PMID 21596538.
- ^ The British Invasion Finding Your Roots
- ^ a b "Mayflower DNA Project". mayflowerdna.org. Retrieved 2020-11-23.
- ^ "FamilyTreeDNA – Lee Surname DNA Research Project (and Leigh, Lea, etc)". familytreedna.com. Retrieved 2020-12-10.
- ^ "FamilyTreeDNA – Grimaldi Genealogy Project at FtDNA". familytreedna.com. Retrieved 2020-12-11.
- ^ "Jackson DNA Project". FamilyTreeDNA. 11 December 2020.
- ^ Hay M (July 2020). "Origins and history of Haplogroup I1 (Y-DNA)". ResearchGate. 1: 9.
- ^ a b c Maciamo E. "Haplogroup I1 (Y-DNA)". Eupedia. Retrieved 2020-12-11.
- ^ Петр Толстой. Моя родословная. Выпуск от 18.04.2010 (in Russian), retrieved 2020-12-15
- ^ "I-BY229 YTree". yfull.com. Retrieved 2020-12-10.
- ^ "Swedenborg". Höijen (in Swedish). Archived from the original on 2020-10-26. Retrieved 2020-12-10.
- ^ "Claas Jansz van Rensburg, SV/PROG". geni_family_tree. Retrieved 2021-01-03.
- ^ "janse /jansen van Rensburg I-M253 genealogy discussion". geni_family_tree. Retrieved 2021-01-03.
- ^ "Rolf H. Nevanlinna". geni_family_tree. Retrieved 2020-12-26.
- ^ olenus (2018-03-30). "I1: Rolf Nevanlinna (né Neovius)". Descendants of haplogroup IJ-M429. Retrieved 2020-12-26.
- ^ "Arne Edvard Nevanlinna". geni_family_tree. Retrieved 2020-12-26.
- ^ "Morse/Moss DNA Testing". morsesociety.org. Retrieved 2020-12-10.
- ^ "FamilyTreeDNA – Morse / Moss DNA Project". familytreedna.com. Retrieved 2020-12-10.
- ^ "Peter Morse's Family Tree". geni.com. Retrieved 2020-12-10.
- ^ "FamilyTreeDNA – Sweden DNA Project – Sverigeprojektet". familytreedna.com. Retrieved 2021-02-14.
- ^ "Eskilstuna kommun · EM GN398 – Familjen Larsson, Torshälla ca 1900". Eskilstuna kommun (in Swedish). Retrieved 2021-02-14.
- ^ "I-Y24470 YTree". yfull.com. Retrieved 2021-02-14.
- ^ "Familjen Larssons Anfäder". hosserudkullen.se. Retrieved 2021-02-14.
- ^ Archived at Ghostarchive and the Wayback Machine: "I Did A DNA Test... (I Guess Im Cancelled Now)" – via YouTube.
- ^ "FamilyTreeDNA – The Norway DNA Project – Norgesprosjektet". familytreedna.com. Retrieved 2021-02-02.
- ^ Tovseth A (June 2018). "Andrésen, Färnskog & Hansen family research". Kjellivar.tripod.com. Retrieved 2 February 2021.
- ^ "Personer med namnet Andresen | Släktingar.se". slaktingar.se. Retrieved 2021-02-02.
- ^ "Björn Andresen: Min passion för mamma blev aldrig besvarad – Katarina Hahr möter". Radio Sveriges (in Swedish). Retrieved 2021-02-02.
- ^ "Johan Peter Andresen – Ancestry". ancestry.se. Retrieved 2021-02-02.
- ^ "Family tree of Daniel Andresen". Geneanet. Retrieved 2021-02-02.
- ^ "FamilyTreeDNA – Pine/Pyne Genealogy DNA Project". familytreedna.com. Retrieved 2020-12-10.
- ^ "James Pine, Sr". geni_family_tree. Retrieved 2020-12-10.
- ^ Janlind F (20 February 2021). "Bianca Salming om relationen med Börje: "Känner mig hemsk"" [Bianca Salming on her relationship with Börje: "Feeling awful"]. Goteborgs-Posten (in Swedish).
- ^ Mercedes (2023-03-26). "Beethoven DNA Discovery – Find Out If You Are Related". Who are You Made Of?. Retrieved 2023-10-07.
- ^ snpdev. "Reference SNP (refSNP) Cluster Report: rs9341296". nih.gov.
- ^ snpdev. "Reference SNP (refSNP) Cluster Report: rs13447354". nih.gov.
- ^ P30[permanent dead link]
- ^ P40[permanent dead link]
Further reading
[edit]- Allentoft ME, Sikora M, Sjögren KG, Rasmussen S, Rasmussen M, Stenderup J, et al. (June 2015). "Population genomics of Bronze Age Eurasia". Nature. 522 (7555): 167–72. Bibcode:2015Natur.522..167A. doi:10.1038/nature14507. PMID 26062507. S2CID 4399103.
- Brunel S, Bennett EA, Cardin L, Garraud D, Barrand Emam H, Beylier A, et al. (June 2020). "Ancient genomes from present-day France unveil 7,000 years of its demographic history". Proceedings of the National Academy of Sciences of the United States of America. 117 (23): 12791–98. Bibcode:2020PNAS..11712791B. doi:10.1073/pnas.1918034117. PMC 7293694. PMID 32457149.
- Malmström H, Gilbert MT, Thomas MG, Brandström M, Storå J, Molnar P, et al. (November 2009). "Ancient DNA reveals lack of continuity between neolithic hunter-gatherers and contemporary Scandinavians". Current Biology. 19 (20): 1758–62. doi:10.1016/j.cub.2009.09.017. PMID 19781941. S2CID 9487217.
- Malmström H, Günther T, Svensson EM, Juras A, Fraser M, Munters AR, Pospieszny Ł, Tõrv M, Lindström J, Götherström A, Storå J, Jakobsson M (October 2019). "The genomic ancestry of the Scandinavian Battle Axe Culture people and their relation to the broader Corded Ware horizon". Proceedings. Biological Sciences. 286 (1912): 20191528. doi:10.1098/rspb.2019.1528. PMC 6790770. PMID 31594508.
- Villalba-Mouco V, van de Loosdrecht MS, Posth C, Mora R, Martínez-Moreno J, Rojo-Guerra M, et al. (April 2019). "Survival of Late Pleistocene Hunter-Gatherer Ancestry in the Iberian Peninsula". Current Biology. 29 (7). Cell Press: 1169–77.e7. doi:10.1016/j.cub.2019.02.006. hdl:10261/208851. PMID 30880015. S2CID 76663708.
External links
[edit]- Haplogroup I databases
- Haplogroup I1 Project at FTDNA
- Danish Demes Regional DNA Project at FTDNA
- Haplogroup I-P109 Project
- British Isles DNA Project
- General Y-DNA databases
There are several public access databases featuring I-M253, including:
- [1] Archived 2011-01-04 at the Wayback Machine
- YHRD : Y-Chromosome STR Haplotype Reference Database
- I1 YTree
