Genome
| Part of a series on |
| Genetics |
|---|
 |

In the fields of molecular biology and genetics, a genome is all the genetic information of an organism.[1] It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of junk DNA with no evident function.[2][3] Almost all eukaryotes have mitochondria and a small mitochondrial genome.[2] Algae and plants also contain chloroplasts with a chloroplast genome.
The study of the genome is called genomics. The genomes of many organisms have been sequenced and various regions have been annotated. The Human Genome Project was started in October 1990, and then reported the sequence of the human genome in April 2003,[4] although the initial "finished" sequence was missing 8% of the genome consisting mostly of repetitive sequences.[5]
With advancements in technology that could handle sequencing of the many repetitive sequences found in human DNA that were not fully uncovered by the original Human Genome Project study, scientists reported the first end-to-end human genome sequence in March 2022.[6]
Origin of the term
[edit]The term genome was created in 1920 by Hans Winkler,[7] professor of botany at the University of Hamburg, Germany. The website Oxford Dictionaries and the Online Etymology Dictionary suggest the name is a blend of the words gene and chromosome.[8][9][10][11] However, see omics for a more thorough discussion. A few related -ome words already existed, such as biome and rhizome, forming a vocabulary into which genome fits systematically.[12]
Definition
[edit]It is very difficult to come up with a precise definition of "genome." It usually refers to the DNA (or sometimes RNA) molecules that carry the genetic information in an organism but sometimes it is difficult to decide which molecules to include in the definition; for example, bacteria usually have one or two large DNA molecules (chromosomes) that contain all of the essential genetic material but they also contain smaller extrachromosomal plasmid molecules that carry important genetic information. The definition of 'genome' that is commonly used in the scientific literature is usually restricted to the large chromosomal DNA molecules in bacteria.[13]
Nuclear genome
[edit]Eukaryotic genomes are even more difficult to define because almost all eukaryotic species contain nuclear chromosomes plus extra DNA molecules in the mitochondria. In addition, algae and plants have chloroplast DNA. Most textbooks make a distinction between the nuclear genome and the organelle (mitochondria and chloroplast) genomes so when they speak of, say, the human genome, they are only referring to the genetic material in the nucleus.[2][14] This is the most common use of 'genome' in the scientific literature.
Ploidy
[edit]Most eukaryotes are diploid, meaning that there are two of each chromosome in the nucleus but the 'genome' refers to only one copy of each chromosome. Some eukaryotes have distinctive sex chromosomes, such as the X and Y chromosomes of mammals, so the technical definition of the genome must include both copies of the sex chromosomes. For example, the standard reference genome of humans consists of one copy of each of the 22 autosomes plus one X chromosome and one Y chromosome.[15]
Sequencing and mapping
[edit]A genome sequence is the complete list of the nucleotides (A, C, G, and T for DNA genomes) that make up all the chromosomes of an individual or a species. Within a species, the vast majority of nucleotides are identical between individuals, but sequencing multiple individuals is necessary to understand the genetic diversity.

In 1976, Walter Fiers at the University of Ghent (Belgium) was the first to establish the complete nucleotide sequence of a viral RNA-genome (Bacteriophage MS2). The next year, Fred Sanger completed the first DNA-genome sequence: Phage Φ-X174, of 5386 base pairs.[16] The first bacterial genome to be sequenced was that of Haemophilus influenzae, completed by a team at The Institute for Genomic Research in 1995. A few months later, the first eukaryotic genome was completed, with sequences of the 16 chromosomes of budding yeast Saccharomyces cerevisiae published as the result of a European-led effort begun in the mid-1980s. The first genome sequence for an archaeon, Methanococcus jannaschii, was completed in 1996, again by The Institute for Genomic Research.[citation needed]
The development of new technologies has made genome sequencing dramatically cheaper and easier, and the number of complete genome sequences is growing rapidly. The US National Institutes of Health maintains one of several comprehensive databases of genomic information.[17] Among the thousands of completed genome sequencing projects include those for rice, a mouse, the plant Arabidopsis thaliana, the puffer fish, and the bacteria E. coli. In December 2013, scientists first sequenced the entire genome of a Neanderthal, an extinct species of humans. The genome was extracted from the toe bone of a 130,000-year-old Neanderthal found in a Siberian cave.[18][19]
New sequencing technologies, such as massive parallel sequencing have also opened up the prospect of personal genome sequencing as a diagnostic tool, as pioneered by Manteia Predictive Medicine. A major step toward that goal was the completion in 2007 of the full genome of James D. Watson, one of the co-discoverers of the structure of DNA.[20]
Whereas a genome sequence lists the order of every DNA base in a genome, a genome map identifies the landmarks. A genome map is less detailed than a genome sequence and aids in navigating around the genome. The Human Genome Project was organized to map and to sequence the human genome. A fundamental step in the project was the release of a detailed genomic map by Jean Weissenbach and his team at the Genoscope in Paris.[21][22]
Reference genome sequences and maps continue to be updated, removing errors and clarifying regions of high allelic complexity.[23] The decreasing cost of genomic mapping has permitted genealogical sites to offer it as a service,[24] to the extent that one may submit one's genome to crowdsourced scientific endeavours such as DNA.LAND at the New York Genome Center,[25] an example both of the economies of scale and of citizen science.[26]
Viral genomes
[edit]Viral genomes can be composed of either RNA or DNA. The genomes of RNA viruses can be either single-stranded RNA or double-stranded RNA, and may contain one or more separate RNA molecules (segments: monopartit or multipartit genome). DNA viruses can have either single-stranded or double-stranded genomes. Most DNA virus genomes are composed of a single, linear molecule of DNA, but some are made up of a circular DNA molecule.[27]
Prokaryotic genomes
[edit]Prokaryotes and eukaryotes have DNA genomes. Archaea and most bacteria have a single circular chromosome,[28] however, some bacterial species have linear or multiple chromosomes.[29][30] If the DNA is replicated faster than the bacterial cells divide, multiple copies of the chromosome can be present in a single cell, and if the cells divide faster than the DNA can be replicated, multiple replication of the chromosome is initiated before the division occurs, allowing daughter cells to inherit complete genomes and already partially replicated chromosomes. Most prokaryotes have very little repetitive DNA in their genomes.[31] However, some symbiotic bacteria (e.g. Serratia symbiotica) have reduced genomes and a high fraction of pseudogenes: only ~40% of their DNA encodes proteins.[32][33]
Some bacteria have auxiliary genetic material, also part of their genome, which is carried in plasmids. For this, the word genome should not be used as a synonym of chromosome.
Eukaryotic genomes
[edit]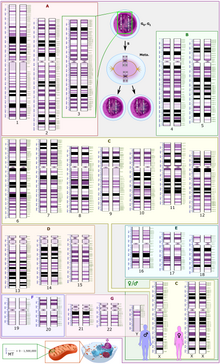
Eukaryotic genomes are composed of one or more linear DNA chromosomes. The number of chromosomes varies widely from Jack jumper ants and an asexual nemotode,[34] which each have only one pair, to a fern species that has 720 pairs.[35] It is surprising the amount of DNA that eukaryotic genomes contain compared to other genomes. The amount is even more than what is necessary for DNA protein-coding and noncoding genes due to the fact that eukaryotic genomes show as much as 64,000-fold variation in their sizes.[36] However, this special characteristic is caused by the presence of repetitive DNA, and transposable elements (TEs).
A typical human cell has two copies of each of 22 autosomes, one inherited from each parent, plus two sex chromosomes, making it diploid. Gametes, such as ova, sperm, spores, and pollen, are haploid, meaning they carry only one copy of each chromosome. In addition to the chromosomes in the nucleus, organelles such as the chloroplasts and mitochondria have their own DNA. Mitochondria are sometimes said to have their own genome often referred to as the "mitochondrial genome". The DNA found within the chloroplast may be referred to as the "plastome". Like the bacteria they originated from, mitochondria and chloroplasts have a circular chromosome.
Unlike prokaryotes where exon-intron organization of protein coding genes exists but is rather exceptional, eukaryotes generally have these features in their genes and their genomes contain variable amounts of repetitive DNA. In mammals and plants, the majority of the genome is composed of repetitive DNA.[37]
DNA sequencing
[edit]High-throughput technology makes sequencing to assemble new genomes accessible to everyone. Sequence polymorphisms are typically discovered by comparing resequenced isolates to a reference, whereas analyses of coverage depth and mapping topology can provide details regarding structural variations such as chromosomal translocations and segmental duplications.
Coding sequences
[edit]DNA sequences that carry the instructions to make proteins are referred to as coding sequences. The proportion of the genome occupied by coding sequences varies widely. A larger genome does not necessarily contain more genes, and the proportion of non-repetitive DNA decreases along with increasing genome size in complex eukaryotes.[37]
Noncoding sequences
[edit]Noncoding sequences include introns, sequences for non-coding RNAs, regulatory regions, and repetitive DNA. Noncoding sequences make up 98% of the human genome. There are two categories of repetitive DNA in the genome: tandem repeats and interspersed repeats.[38]
Tandem repeats
[edit]Short, non-coding sequences that are repeated head-to-tail are called tandem repeats. Microsatellites consisting of 2–5 basepair repeats, while minisatellite repeats are 30–35 bp. Tandem repeats make up about 4% of the human genome and 9% of the fruit fly genome.[39] Tandem repeats can be functional. For example, telomeres are composed of the tandem repeat TTAGGG in mammals, and they play an important role in protecting the ends of the chromosome.
In other cases, expansions in the number of tandem repeats in exons or introns can cause disease.[40] For example, the human gene huntingtin (Htt) typically contains 6–29 tandem repeats of the nucleotides CAG (encoding a polyglutamine tract). An expansion to over 36 repeats results in Huntington's disease, a neurodegenerative disease. Twenty human disorders are known to result from similar tandem repeat expansions in various genes. The mechanism by which proteins with expanded polygulatamine tracts cause death of neurons is not fully understood. One possibility is that the proteins fail to fold properly and avoid degradation, instead accumulating in aggregates that also sequester important transcription factors, thereby altering gene expression.[40]
Tandem repeats are usually caused by slippage during replication, unequal crossing-over and gene conversion.[41]
Transposable elements
[edit]Transposable elements (TEs) are sequences of DNA with a defined structure that are able to change their location in the genome.[39][31][42] TEs are categorized as either as a mechanism that replicates by copy-and-paste or as a mechanism that can be excised from the genome and inserted at a new location. In the human genome, there are three important classes of TEs that make up more than 45% of the human DNA; these classes are The long interspersed nuclear elements (LINEs), The interspersed nuclear elements (SINEs), and endogenous retroviruses. These elements have a big potential to modify the genetic control in a host organism.[36]
The movement of TEs is a driving force of genome evolution in eukaryotes because their insertion can disrupt gene functions, homologous recombination between TEs can produce duplications, and TE can shuffle exons and regulatory sequences to new locations.[43]
Retrotransposons
[edit]Retrotransposons[44] are found mostly in eukaryotes but not found in prokaryotes. Retrotransposons form a large portion of the genomes of many eukaryotes. A retrotransposon is a transposable element that transposes through an RNA intermediate. Retrotransposons[45] are composed of DNA, but are transcribed into RNA for transposition, then the RNA transcript is copied back to DNA formation with the help of a specific enzyme called reverse transcriptase. A retrotransposon that carries reverse transcriptase in its sequence can trigger its own transposition but retrotransposons that lack a reverse transcriptase must use reverse transcriptase synthesized by another retrotransposon. Retrotransposons can be transcribed into RNA, which are then duplicated at another site into the genome.[46] Retrotransposons can be divided into long terminal repeats (LTRs) and non-long terminal repeats (Non-LTRs).[43]
Long terminal repeats (LTRs) are derived from ancient retroviral infections, so they encode proteins related to retroviral proteins including gag (structural proteins of the virus), pol (reverse transcriptase and integrase), pro (protease), and in some cases env (envelope) genes.[42] These genes are flanked by long repeats at both 5' and 3' ends. It has been reported that LTRs consist of the largest fraction in most plant genome and might account for the huge variation in genome size.[47]
Non-long terminal repeats (Non-LTRs) are classified as long interspersed nuclear elements (LINEs), short interspersed nuclear elements (SINEs), and Penelope-like elements (PLEs). In Dictyostelium discoideum, there is another DIRS-like elements belong to Non-LTRs. Non-LTRs are widely spread in eukaryotic genomes.[48]
Long interspersed elements (LINEs) encode genes for reverse transcriptase and endonuclease, making them autonomous transposable elements. The human genome has around 500,000 LINEs, taking around 17% of the genome.[49]
Short interspersed elements (SINEs) are usually less than 500 base pairs and are non-autonomous, so they rely on the proteins encoded by LINEs for transposition.[50] The Alu element is the most common SINE found in primates. It is about 350 base pairs and occupies about 11% of the human genome with around 1,500,000 copies.[43]
DNA transposons
[edit]DNA transposons encode a transposase enzyme between inverted terminal repeats. When expressed, the transposase recognizes the terminal inverted repeats that flank the transposon and catalyzes its excision and reinsertion in a new site.[39] This cut-and-paste mechanism typically reinserts transposons near their original location (within 100 kb).[43] DNA transposons are found in bacteria and make up 3% of the human genome and 12% of the genome of the roundworm C. elegans.[43]
Genome size
[edit]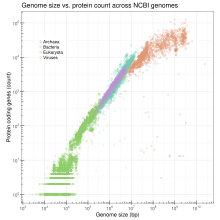
Genome size is the total number of the DNA base pairs in one copy of a haploid genome. Genome size varies widely across species. Invertebrates have small genomes, this is also correlated to a small number of transposable elements. Fish and Amphibians have intermediate-size genomes, and birds have relatively small genomes but it has been suggested that birds lost a substantial portion of their genomes during the phase of transition to flight. Before this loss, DNA methylation allows the adequate expansion of the genome.[36]
In humans, the nuclear genome comprises approximately 3.1 billion nucleotides of DNA, divided into 24 linear molecules, the shortest 45 000 000 nucleotides in length and the longest 248 000 000 nucleotides, each contained in a different chromosome.[51] There is no clear and consistent correlation between morphological complexity and genome size in either prokaryotes or lower eukaryotes.[37][52] Genome size is largely a function of the expansion and contraction of repetitive DNA elements.
Since genomes are very complex, one research strategy is to reduce the number of genes in a genome to the bare minimum and still have the organism in question survive. There is experimental work being done on minimal genomes for single cell organisms as well as minimal genomes for multi-cellular organisms (see developmental biology). The work is both in vivo and in silico.[53][54]
Genome size differences due to transposable elements
[edit]
There are many enormous differences in size in genomes, specially mentioned before in the multicellular eukaryotic genomes. Much of this is due to the differing abundances of transposable elements, which evolve by creating new copies of themselves in the chromosomes.[36] Eukaryote genomes often contain many thousands of copies of these elements, most of which have acquired mutations that make them defective. Here is a table of some significant or representative genomes. See #See also for lists of sequenced genomes.
| Organism type | Organism | Genome size (base pairs) |
Approx. no. of genes | Note | |
|---|---|---|---|---|---|
| Virus | Porcine circovirus type 1 | 1,759 | 1.8 kB | Smallest viruses replicating autonomously in eukaryotic cells[55] | |
| Virus | Bacteriophage MS2 | 3,569 | 3.6 kB | First sequenced RNA-genome[56] | |
| Virus | SV40 | 5,224 | 5.2 kB | [57] | |
| Virus | Phage Φ-X174 | 5,386 | 5.4 kB | First sequenced DNA-genome[58] | |
| Virus | HIV | 9,749 | 9.7 kB | [59] | |
| Virus | Phage λ | 48,502 | 48.5 kB | Often used as a vector for the cloning of recombinant DNA | |
| Virus | Megavirus | 1,259,197 | 1.3 MB | Until 2013 the largest known viral genome[63] | |
| Virus | Pandoravirus salinus | 2,470,000 | 2.47 MB | Largest known viral genome.[64] | |
| Eukaryotic organelle | Human mitochondrion | 16,569 | 16.6 kB | [65] | |
| Bacterium | Nasuia deltocephalinicola (strain NAS-ALF) | 112,091 | 112 kB | 137 | Smallest known non-viral genome. Symbiont of leafhoppers.[66] |
| Bacterium | Carsonella ruddii | 159,662 | 160 kB | An endosymbiont of psyllid insects | |
| Bacterium | Buchnera aphidicola | 600,000 | 600 kB | An endosymbiont of aphids[67] | |
| Bacterium | Wigglesworthia glossinidia | 700,000 | 700 kB | A symbiont in the gut of the tsetse fly | |
| Bacterium – cyanobacterium | Prochlorococcus spp. (1.7 Mb) | 1,700,000 | 1.7 MB | 1,884 | Smallest known cyanobacterium genome. One of the primary photosynthesizers on Earth.[68][69] |
| Bacterium | Haemophilus influenzae | 1,830,000 | 1.8 MB | First genome of a living organism sequenced, July 1995[70] | |
| Bacterium | Escherichia coli | 4,600,000 | 4.6 MB | 4,288 | [71] |
| Bacterium – cyanobacterium | Nostoc punctiforme | 9,000,000 | 9 MB | 7,432 | 7432 open reading frames[72] |
| Bacterium | Solibacter usitatus (strain Ellin 6076) | 9,970,000 | 10 MB | [73] | |
| Bacterium | Sorangium cellulosum | 13,033,779 | 13 MB | 9,367 | Largest known bacterial genome[74][75] |
| Amoeboid | Polychaos dubium ("Amoeba" dubia) | 670,000,000,000 | 670 GB | Largest known genome.[76] (Disputed)[77] | |
| Plant | Genlisea tuberosa | 61,000,000 | 61 MB | Smallest recorded flowering plant genome, 2014[78] | |
| Plant | Arabidopsis thaliana | 135,000,000[79] | 135 MB | 27,655[80] | First plant genome sequenced, December 2000[81] |
| Plant | Populus trichocarpa | 480,000,000 | 480 MB | 73,013 | First tree genome sequenced, September 2006[82] |
| Plant | Pinus taeda (Loblolly pine) | 22,180,000,000 | 22.18 GB | 50,172 | Gymnosperms generally have much larger genomes than angiosperms[83][84] |
| Plant | Fritillaria assyriaca | 130,000,000,000 | 130 GB | ||
| Plant | Paris japonica (Japanese-native, order Liliales) | 150,000,000,000 | 150 GB | Former largest plant genome known[85] | |
| Plant – moss | Physcomitrella patens | 480,000,000 | 480 MB | First genome of a bryophyte sequenced, January 2008[86] | |
| Fungus – yeast | Saccharomyces cerevisiae | 12,100,000 | 12.1 MB | 6,294 | First eukaryotic genome sequenced, 1996[87] |
| Fungus | Aspergillus nidulans | 30,000,000 | 30 MB | 9,541 | [88] |
| Nematode | Pratylenchus coffeae | 20,000,000 | 20 MB | [89] Smallest animal genome known[90] | |
| Nematode | Caenorhabditis elegans | 100,300,000 | 100 MB | 19,000 | First multicellular animal genome sequenced, December 1998[91] |
| Insect | Belgica antarctica (Antarctic midge) | 99,000,000 | 99 MB | Smallest insect genome sequenced thus far, likely an adaptation to an extreme environment[92] | |
| Insect | Drosophila melanogaster (fruit fly) | 175,000,000 | 175 MB | 13,600 | Size variation based on strain (175–180 Mb; standard y w strain is 175 Mb)[93] |
| Insect | Apis mellifera (honey bee) | 236,000,000 | 236 MB | 10,157 | [94] |
| Insect | Bombyx mori (silk moth) | 432,000,000 | 432 MB | 14,623 | 14,623 predicted genes[95] |
| Insect | Solenopsis invicta (fire ant) | 480,000,000 | 480 MB | 16,569 | [96] |
| Crustacean | Antarctic krill | 48,010,000,000 | 48 GB | 23,000 | 70-92% repetitive DNA[97] |
| Amphibian | Neuse River waterdog | 118,000,000,000 | 118 GB | Largest tetrapod genome sequenced as of 2022[98] | |
| Amphibian | Ornate burrowing frog | 1,060,000,000 | 1.06 GB | Smallest known frog genome[99] | |
| Lizard | Armadillo girdled lizard | 3,930,000,000 | 3.93 GB | Largest known squamate genome[100] | |
| Mammal | Plains viscacha rat | 8,400,000,000 | 8.4 GB | Largest known mammalian genome[101] | |
| Mammal | Mus musculus | 2,700,000,000 | 2.7 GB | 20,210 | [102] |
| Mammal | Pan paniscus | 3,286,640,000 | 3.3 GB | 20,000 | Bonobo – estimated genome size 3.29 billion bp[103] |
| Mammal | Homo sapiens | 3,117,000,000 | 3.1 GB | 20,000 | Homo sapiens genome size estimated at 3.12 Gbp in 2022[51]
Initial sequencing and analysis of the human genome[104] |
| Bird | Gallus gallus | 1,043,000,000 | 1.0 GB | 20,000 | [105] |
| Fish | Tetraodon nigroviridis (type of puffer fish) | 385,000,000 | 390 MB | Smallest vertebrate genome known, estimated to be 340 Mb[106][107] – 385 Mb[108] | |
| Fish | Protopterus aethiopicus (marbled lungfish) | 130,000,000,000 | 130 GB | Largest vertebrate genome known[109][110][111] | |
| Plant | Tmesipteris truncata (fern ally) | 160,000,000,000 | 160 GB | Largest plant genome known[112] | |
Genomic alterations
[edit]All the cells of an organism originate from a single cell, so they are expected to have identical genomes; however, in some cases, differences arise. Both the process of copying DNA during cell division and exposure to environmental mutagens can result in mutations in somatic cells. In some cases, such mutations lead to cancer because they cause cells to divide more quickly and invade surrounding tissues.[113] In certain lymphocytes in the human immune system, V(D)J recombination generates different genomic sequences such that each cell produces a unique antibody or T cell receptors.
During meiosis, diploid cells divide twice to produce haploid germ cells. During this process, recombination results in a reshuffling of the genetic material from homologous chromosomes so each gamete has a unique genome.
Genome-wide reprogramming
[edit]Genome-wide reprogramming in mouse primordial germ cells involves epigenetic imprint erasure leading to totipotency. Reprogramming is facilitated by active DNA demethylation, a process that entails the DNA base excision repair pathway.[114] This pathway is employed in the erasure of CpG methylation (5mC) in primordial germ cells. The erasure of 5mC occurs via its conversion to 5-hydroxymethylcytosine (5hmC) driven by high levels of the ten-eleven dioxygenase enzymes TET1 and TET2.[115]
Genome evolution
[edit]Genomes are more than the sum of an organism's genes and have traits that may be measured and studied without reference to the details of any particular genes and their products. Researchers compare traits such as karyotype (chromosome number), genome size, gene order, codon usage bias, and GC-content to determine what mechanisms could have produced the great variety of genomes that exist today (for recent overviews, see Brown 2002; Saccone and Pesole 2003; Benfey and Protopapas 2004; Gibson and Muse 2004; Reese 2004; Gregory 2005).
Duplications play a major role in shaping the genome. Duplication may range from extension of short tandem repeats, to duplication of a cluster of genes, and all the way to duplication of entire chromosomes or even entire genomes. Such duplications are probably fundamental to the creation of genetic novelty.
Horizontal gene transfer is invoked to explain how there is often an extreme similarity between small portions of the genomes of two organisms that are otherwise very distantly related. Horizontal gene transfer seems to be common among many microbes. Also, eukaryotic cells seem to have experienced a transfer of some genetic material from their chloroplast and mitochondrial genomes to their nuclear chromosomes. Recent empirical data suggest an important role of viruses and sub-viral RNA-networks to represent a main driving role to generate genetic novelty and natural genome editing.
In fiction
[edit]Works of science fiction illustrate concerns about the availability of genome sequences.
Michael Crichton's 1990 novel Jurassic Park and the subsequent film tell the story of a billionaire who creates a theme park of cloned dinosaurs on a remote island, with disastrous outcomes. A geneticist extracts dinosaur DNA from the blood of ancient mosquitoes and fills in the gaps with DNA from modern species to create several species of dinosaurs. A chaos theorist is asked to give his expert opinion on the safety of engineering an ecosystem with the dinosaurs, and he repeatedly warns that the outcomes of the project will be unpredictable and ultimately uncontrollable. These warnings about the perils of using genomic information are a major theme of the book.
The 1997 film Gattaca is set in a futurist society where genomes of children are engineered to contain the most ideal combination of their parents' traits, and metrics such as risk of heart disease and predicted life expectancy are documented for each person based on their genome. People conceived outside of the eugenics program, known as "In-Valids" suffer discrimination and are relegated to menial occupations. The protagonist of the film is an In-Valid who works to defy the supposed genetic odds and achieve his dream of working as a space navigator. The film warns against a future where genomic information fuels prejudice and extreme class differences between those who can and cannot afford genetically engineered children.[116]
See also
[edit]- Bacterial genome size
- Cryoconservation of animal genetic resources
- Genome Browser
- Genome Compiler
- Genome topology
- Genome-wide association study
- List of sequenced animal genomes
- List of sequenced archaeal genomes
- List of sequenced bacterial genomes
- List of sequenced eukaryotic genomes
- List of sequenced fungi genomes
- List of sequenced plant genomes
- List of sequenced plastomes
- List of sequenced protist genomes
- Metagenomics
- Microbiome
- Molecular epidemiology
- Molecular pathological epidemiology
- Molecular pathology
- Nucleic acid sequence
- Pan-genome
- Precision medicine
- Regulator gene
- Whole genome sequencing
References
[edit]- ^ Roth, Stephanie Clare (1 July 2019). "What is genomic medicine?". Journal of the Medical Library Association. 107 (3). University Library System, University of Pittsburgh: 442–448. doi:10.5195/jmla.2019.604. ISSN 1558-9439. PMC 6579593. PMID 31258451.
- ^ a b c Graur, Dan; Sater, Amy K.; Cooper, Tim F. (2016). Molecular and Genome Evolution. Sinauer Associates, Inc. ISBN 9781605354699. OCLC 951474209.
- ^ Brosius, J (2009). "The Fragmented Gene". Annals of the New York Academy of Sciences. 1178 (1): 186–93. Bibcode:2009NYASA1178..186B. doi:10.1111/j.1749-6632.2009.05004.x. PMID 19845638. S2CID 8279434.
- ^ "The Human Genome Project". Genome.gov. Retrieved 29 April 2023.
- ^ "First complete sequence of a human genome". National Institutes of Health (NIH). 11 April 2022. Archived from the original on 14 April 2023. Retrieved 29 April 2023.
- ^ Hartley, Gabrielle (31 March 2022). "The Human Genome Project pieced together only 92% of the DNA – now scientists have finally filled in the remaining 8%". TheConversation.org. The Conversation US, Inc. Retrieved 4 April 2022.
- ^ Winkler HL (1920). Verbreitung und Ursache der Parthenogenesis im Pflanzen- und Tierreiche. Jena: Verlag Fischer.
- ^ "definition of Genome in Oxford dictionary". Archived from the original on 1 March 2014. Retrieved 25 March 2014.
- ^ "genome". Oxford English Dictionary (Online ed.). Oxford University Press. (Subscription or participating institution membership required.)
- ^ "genome". Lexico UK English Dictionary. Oxford University Press. Archived from the original on 24 August 2022.
- ^ Harper, Douglas. "genome". Online Etymology Dictionary.
- ^ Lederberg J, McCray AT (2001). "'Ome Sweet 'Omics – A Genealogical Treasury of Words" (PDF). The Scientist. 15 (7). Archived from the original (PDF) on 29 September 2006.
- ^ Kirchberger PC, Schmidt ML, and Ochman H (2020). "The ingenuity of bacterial genomes". Annual Review of Microbiology. 74: 815–834. doi:10.1146/annurev-micro-020518-115822. PMID 32692614. S2CID 220699395.
- ^ Brown, TA (2018). Genomes 4. New York, NY, USA: Garland Science. ISBN 9780815345084.
- ^ "Ensembl Human Assembly and gene annotation (GRCh38)". Ensembl. Retrieved 30 May 2022.
- ^ "All about genes". beowulf.org.uk.
- ^ "Genome Home". 8 December 2010. Retrieved 27 January 2011.
- ^ Zimmer C (18 December 2013). "Toe Fossil Provides Complete Neanderthal Genome". The New York Times. Archived from the original on 2 January 2022. Retrieved 18 December 2013.
- ^ Prüfer K, Racimo F, Patterson N, Jay F, Sankararaman S, Sawyer S, et al. (January 2014). "The complete genome sequence of a Neanderthal from the Altai Mountains". Nature. 505 (7481): 43–49. Bibcode:2014Natur.505...43P. doi:10.1038/nature12886. PMC 4031459. PMID 24352235.
- ^ Wade N (31 May 2007). "Genome of DNA Pioneer Is Deciphered". The New York Times. Retrieved 2 April 2010.
- ^ "What's a Genome?". Genomenewsnetwork.org. 15 January 2003. Retrieved 27 January 2011.
- ^ "Mapping Factsheet". 29 March 2004. Archived from the original on 19 July 2010. Retrieved 27 January 2011.
- ^ Genome Reference Consortium. "Assembling the Genome". Retrieved 23 August 2016.
- ^ Kaplan, Sarah (17 April 2016). "How do your 20,000 genes determine so many wildly different traits? They multitask". The Washington Post. Retrieved 27 August 2016.
- ^ Check Hayden, Erika (2015). "Scientists hope to attract millions to 'DNA.LAND'". Nature. doi:10.1038/nature.2015.18514. S2CID 211729308.
- ^ Zimmer, Carl (25 July 2016). "Game of Genomes, Episode 13: Answers and Questions". STAT. Retrieved 27 August 2016.
- ^ Gelderblom, Hans R. (1996). Structure and Classification of Viruses (4th ed.). Galveston, TX: The University of Texas Medical Branch at Galveston. ISBN 9780963117212. PMID 21413309.
- ^ Samson RY, Bell SD (2014). "Archaeal chromosome biology". Journal of Molecular Microbiology and Biotechnology. 24 (5–6): 420–27. doi:10.1159/000368854. PMC 5175462. PMID 25732343.
- ^ Chaconas G, Chen CW (2005). "Replication of Linear Bacterial Chromosomes: No Longer Going Around in Circles". The Bacterial Chromosome. pp. 525–540. doi:10.1128/9781555817640.ch29. ISBN 9781555812324.
- ^ "Bacterial Chromosomes". Microbial Genetics. 2002.
- ^ a b Koonin EV, Wolf YI (July 2010). "Constraints and plasticity in genome and molecular-phenome evolution". Nature Reviews. Genetics. 11 (7): 487–98. doi:10.1038/nrg2810. PMC 3273317. PMID 20548290.
- ^ McCutcheon JP, Moran NA (November 2011). "Extreme genome reduction in symbiotic bacteria". Nature Reviews. Microbiology. 10 (1): 13–26. doi:10.1038/nrmicro2670. PMID 22064560. S2CID 7175976.
- ^ Land M, Hauser L, Jun SR, Nookaew I, Leuze MR, Ahn TH, Karpinets T, Lund O, Kora G, Wassenaar T, Poudel S, Ussery DW (March 2015). "Insights from 20 years of bacterial genome sequencing". Functional & Integrative Genomics. 15 (2): 141–61. doi:10.1007/s10142-015-0433-4. PMC 4361730. PMID 25722247.
- ^ "Scientists sequence asexual tiny worm whose lineage stretches back 18 million years". ScienceDaily. Retrieved 7 November 2017.
- ^ Khandelwal S (March 1990). "Chromosome evolution in the genus Ophioglossum L.". Botanical Journal of the Linnean Society. 102 (3): 205–17. doi:10.1111/j.1095-8339.1990.tb01876.x.
- ^ a b c d Zhou, Wanding; Liang, Gangning; Molloy, Peter L.; Jones, Peter A. (11 August 2020). "DNA methylation enables transposable element-driven genome expansion". Proceedings of the National Academy of Sciences of the United States of America. 117 (32): 19359–19366. Bibcode:2020PNAS..11719359Z. doi:10.1073/pnas.1921719117. ISSN 1091-6490. PMC 7431005. PMID 32719115.
- ^ a b c Lewin B (2004). Genes VIII (8th ed.). Upper Saddle River, NJ: Pearson/Prentice Hall. ISBN 978-0-13-143981-8.
- ^ Stojanovic N, ed. (2007). Computational genomics : current methods. Wymondham: Horizon Bioscience. ISBN 978-1-904933-30-4.
- ^ a b c Padeken J, Zeller P, Gasser SM (April 2015). "Repeat DNA in genome organization and stability". Current Opinion in Genetics & Development. 31: 12–19. doi:10.1016/j.gde.2015.03.009. PMID 25917896.
- ^ a b Usdin K (July 2008). "The biological effects of simple tandem repeats: lessons from the repeat expansion diseases". Genome Research. 18 (7): 1011–19. doi:10.1101/gr.070409.107. PMC 3960014. PMID 18593815.
- ^ Li YC, Korol AB, Fahima T, Beiles A, Nevo E (December 2002). "Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review". Molecular Ecology. 11 (12): 2453–65. Bibcode:2002MolEc..11.2453L. doi:10.1046/j.1365-294X.2002.01643.x. PMID 12453231. S2CID 23606208.
- ^ a b Wessler SR (November 2006). "Transposable elements and the evolution of eukaryotic genomes". Proceedings of the National Academy of Sciences of the United States of America. 103 (47): 17600–01. Bibcode:2006PNAS..10317600W. doi:10.1073/pnas.0607612103. PMC 1693792. PMID 17101965.
- ^ a b c d e Kazazian HH (March 2004). "Mobile elements: drivers of genome evolution". Science. 303 (5664): 1626–32. Bibcode:2004Sci...303.1626K. doi:10.1126/science.1089670. PMID 15016989. S2CID 1956932.
- ^ "Transposon | genetics". Encyclopedia Britannica. Retrieved 5 December 2020.
- ^ Sanders, Mark Frederick (2019). Genetic Analysis: an integrated approach third edition. New York: Pearson, always learning, and mastering. p. 425. ISBN 9780134605173.
- ^ Deininger PL, Moran JV, Batzer MA, Kazazian HH (December 2003). "Mobile elements and mammalian genome evolution". Current Opinion in Genetics & Development. 13 (6): 651–58. doi:10.1016/j.gde.2003.10.013. PMID 14638329.
- ^ Kidwell MG, Lisch DR (March 2000). "Transposable elements and host genome evolution". Trends in Ecology & Evolution. 15 (3): 95–99. doi:10.1016/S0169-5347(99)01817-0. PMID 10675923.
- ^ Richard GF, Kerrest A, Dujon B (December 2008). "Comparative genomics and molecular dynamics of DNA repeats in eukaryotes". Microbiology and Molecular Biology Reviews. 72 (4): 686–727. doi:10.1128/MMBR.00011-08. PMC 2593564. PMID 19052325.
- ^ Cordaux R, Batzer MA (October 2009). "The impact of retrotransposons on human genome evolution". Nature Reviews. Genetics. 10 (10): 691–703. doi:10.1038/nrg2640. PMC 2884099. PMID 19763152.
- ^ Han JS, Boeke JD (August 2005). "LINE-1 retrotransposons: modulators of quantity and quality of mammalian gene expression?". BioEssays. 27 (8): 775–84. doi:10.1002/bies.20257. PMID 16015595. S2CID 26424042.
- ^ a b Nurk, Sergey; et al. (31 March 2022). "The complete sequence of a human genome" (PDF). Science. 376 (6588): 44–53. Bibcode:2022Sci...376...44N. doi:10.1126/science.abj6987. PMC 9186530. PMID 35357919. S2CID 235233625. Archived (PDF) from the original on 26 May 2022.
- ^ Gregory TR, Nicol JA, Tamm H, Kullman B, Kullman K, Leitch IJ, Murray BG, Kapraun DF, Greilhuber J, Bennett MD (January 2007). "Eukaryotic genome size databases". Nucleic Acids Research. 35 (Database issue): D332–38. doi:10.1093/nar/gkl828. PMC 1669731. PMID 17090588.
- ^ Glass JI, Assad-Garcia N, Alperovich N, Yooseph S, Lewis MR, Maruf M, Hutchison CA, Smith HO, Venter JC (January 2006). "Essential genes of a minimal bacterium". Proceedings of the National Academy of Sciences of the United States of America. 103 (2): 425–30. Bibcode:2006PNAS..103..425G. doi:10.1073/pnas.0510013103. PMC 1324956. PMID 16407165.
- ^ Forster AC, Church GM (2006). "Towards synthesis of a minimal cell". Molecular Systems Biology. 2 (1): 45. doi:10.1038/msb4100090. PMC 1681520. PMID 16924266.
- ^ Mankertz P (2008). "Molecular Biology of Porcine Circoviruses". Animal Viruses: Molecular Biology. Caister Academic Press. ISBN 978-1-904455-22-6.
- ^ Fiers W, Contreras R, Duerinck F, Haegeman G, Iserentant D, Merregaert J, Min Jou W, Molemans F, Raeymaekers A, Van den Berghe A, Volckaert G, Ysebaert M (April 1976). "Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene". Nature. 260 (5551): 500–07. Bibcode:1976Natur.260..500F. doi:10.1038/260500a0. PMID 1264203. S2CID 4289674.
- ^ Fiers W, Contreras R, Haegemann G, Rogiers R, Van de Voorde A, Van Heuverswyn H, Van Herreweghe J, Volckaert G, Ysebaert M (May 1978). "Complete nucleotide sequence of SV40 DNA". Nature. 273 (5658): 113–20. Bibcode:1978Natur.273..113F. doi:10.1038/273113a0. PMID 205802. S2CID 1634424.
- ^ Sanger F, Air GM, Barrell BG, Brown NL, Coulson AR, Fiddes CA, Hutchison CA, Slocombe PM, Smith M (February 1977). "Nucleotide sequence of bacteriophage phi X174 DNA". Nature. 265 (5596): 687–95. Bibcode:1977Natur.265..687S. doi:10.1038/265687a0. PMID 870828. S2CID 4206886.
- ^ "Virology – Human Immunodeficiency Virus And Aids, Structure: The Genome And Proteins of HIV". Pathmicro.med.sc.edu. 1 July 2010. Retrieved 27 January 2011.
- ^ Thomason L, Court DL, Bubunenko M, Costantino N, Wilson H, Datta S, Oppenheim A (April 2007). "Recombineering: Genetic Engineering in Bacteria Using Homologous Recombination". Current Protocols in Molecular Biology. Chapter 1: Unit 1.16. doi:10.1002/0471142727.mb0116s78. ISBN 978-0-471-14272-0. PMID 18265390. S2CID 490362.
- ^ Court DL, Oppenheim AB, Adhya SL (January 2007). "A new look at bacteriophage lambda genetic networks". Journal of Bacteriology. 189 (2): 298–304. doi:10.1128/JB.01215-06. PMC 1797383. PMID 17085553.
- ^ Sanger F, Coulson AR, Hong GF, Hill DF, Petersen GB (December 1982). "Nucleotide sequence of bacteriophage lambda DNA". Journal of Molecular Biology. 162 (4): 729–73. doi:10.1016/0022-2836(82)90546-0. PMID 6221115.
- ^ Legendre M, Arslan D, Abergel C, Claverie JM (January 2012). "Genomics of Megavirus and the elusive fourth domain of Life". Communicative & Integrative Biology. 5 (1): 102–06. doi:10.4161/cib.18624. PMC 3291303. PMID 22482024.
- ^ Philippe N, Legendre M, Doutre G, Couté Y, Poirot O, Lescot M, Arslan D, Seltzer V, Bertaux L, Bruley C, Garin J, Claverie JM, Abergel C (July 2013). "Pandoraviruses: amoeba viruses with genomes up to 2.5 Mb reaching that of parasitic eukaryotes" (PDF). Science. 341 (6143): 281–86. Bibcode:2013Sci...341..281P. doi:10.1126/science.1239181. PMID 23869018. S2CID 16877147.
- ^ Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJ, Staden R, Young IG (April 1981). "Sequence and organization of the human mitochondrial genome". Nature. 290 (5806): 457–65. Bibcode:1981Natur.290..457A. doi:10.1038/290457a0. PMID 7219534. S2CID 4355527.
- ^ Bennett GM, Moran NA (5 August 2013). "Small, smaller, smallest: the origins and evolution of ancient dual symbioses in a Phloem-feeding insect". Genome Biology and Evolution. 5 (9): 1675–88. doi:10.1093/gbe/evt118. PMC 3787670. PMID 23918810.
- ^ Shigenobu S, Watanabe H, Hattori M, Sakaki Y, Ishikawa H (September 2000). "Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS". Nature. 407 (6800): 81–86. Bibcode:2000Natur.407...81S. doi:10.1038/35024074. PMID 10993077.
- ^ Rocap G, Larimer FW, Lamerdin J, Malfatti S, Chain P, Ahlgren NA, et al. (August 2003). "Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation". Nature. 424 (6952): 1042–47. Bibcode:2003Natur.424.1042R. doi:10.1038/nature01947. PMID 12917642. S2CID 4344597.
- ^ Dufresne A, Salanoubat M, Partensky F, Artiguenave F, Axmann IM, Barbe V, et al. (August 2003). "Genome sequence of the cyanobacterium Prochlorococcus marinus SS120, a nearly minimal oxyphototrophic genome". Proceedings of the National Academy of Sciences of the United States of America. 100 (17): 10020–25. Bibcode:2003PNAS..10010020D. doi:10.1073/pnas.1733211100. PMC 187748. PMID 12917486.
- ^ Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Bult CJ, Tomb JF, Dougherty BA, Merrick JM (July 1995). "Whole-genome random sequencing and assembly of Haemophilus influenzae Rd". Science. 269 (5223): 496–512. Bibcode:1995Sci...269..496F. doi:10.1126/science.7542800. PMID 7542800. S2CID 10423613.
- ^ Blattner FR, Plunkett G, Bloch CA, Perna NT, Burland V, Riley M, et al. (September 1997). "The complete genome sequence of Escherichia coli K-12". Science. 277 (5331): 1453–62. doi:10.1126/science.277.5331.1453. PMID 9278503.
- ^ Meeks JC, Elhai J, Thiel T, Potts M, Larimer F, Lamerdin J, Predki P, Atlas R (2001). "An overview of the genome of Nostoc punctiforme, a multicellular, symbiotic cyanobacterium". Photosynthesis Research. 70 (1): 85–106. doi:10.1023/A:1013840025518. PMID 16228364. S2CID 8752382.
- ^ Challacombe JF, Eichorst SA, Hauser L, Land M, Xie G, Kuske CR (15 September 2011). Steinke D (ed.). "Biological consequences of ancient gene acquisition and duplication in the large genome of Candidatus Solibacter usitatus Ellin6076". PLOS ONE. 6 (9): e24882. Bibcode:2011PLoSO...624882C. doi:10.1371/journal.pone.0024882. PMC 3174227. PMID 21949776.
- ^ Schneiker, Susanne; Perlova, Olena; Kaiser, Olaf; Gerth, Klaus; Alici, Aysel; Altmeyer, Matthias O; Bartels, Daniela; Bekel, Thomas; Beyer, Stefan; Bode, Edna; Bode, Helge B; Bolten, Christoph J; Choudhuri, Jomuna V; Doss, Sabrina; Elnakady, Yasser A (November 2007). "Complete genome sequence of the myxobacterium Sorangium cellulosum". Nature Biotechnology. 25 (11): 1281–1289. doi:10.1038/nbt1354. ISSN 1087-0156.
- ^ "Largest known bacterial genome size - Bacteria Sorangium cellulosum - BNID 104469". bionumbers.hms.harvard.edu. Retrieved 2 May 2024.
- ^ Parfrey LW, Lahr DJ, Katz LA (April 2008). "The dynamic nature of eukaryotic genomes". Molecular Biology and Evolution. 25 (4): 787–94. doi:10.1093/molbev/msn032. PMC 2933061. PMID 18258610.
- ^ ScienceShot: Biggest Genome Ever Archived 11 October 2010 at the Wayback Machine, comments: "The measurement for Amoeba dubia and other protozoa which have been reported to have very large genomes were made in the 1960s using a rough biochemical approach which is now considered to be an unreliable method for accurate genome size determinations."
- ^ Fleischmann A, Michael TP, Rivadavia F, Sousa A, Wang W, Temsch EM, Greilhuber J, Müller KF, Heubl G (December 2014). "Evolution of genome size and chromosome number in the carnivorous plant genus Genlisea (Lentibulariaceae), with a new estimate of the minimum genome size in angiosperms". Annals of Botany. 114 (8): 1651–63. doi:10.1093/aob/mcu189. PMC 4649684. PMID 25274549.
- ^ "Genome Assembly". The Arabidopsis Information Resource (TAIR).
- ^ "Details - Arabidopsis thaliana - Ensembl Genomes 40". plants.ensembl.org.
- ^ Greilhuber J, Borsch T, Müller K, Worberg A, Porembski S, Barthlott W (November 2006). "Smallest angiosperm genomes found in lentibulariaceae, with chromosomes of bacterial size". Plant Biology. 8 (6): 770–77. Bibcode:2006PlBio...8..770G. doi:10.1055/s-2006-924101. PMID 17203433. S2CID 260252929.
- ^ Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, et al. (September 2006). "The genome of black cottonwood, Populus trichocarpa (Torr. & Gray)" (PDF). Science. 313 (5793): 1596–604. Bibcode:2006Sci...313.1596T. doi:10.1126/science.1128691. OSTI 901819. PMID 16973872. S2CID 7717980.
- ^ Zimin, Aleksey; Stevens, Kristian; et al. (March 2014). "Sequencing and Assembly of the 22-Gb Loblolly Pine Genome". Genetics. 196 (3): 875–890. doi:10.1534/genetics.113.159715. PMC 3948813. PMID 24653210.
- ^ Neale, David B; et al. (March 2014). "Decoding the massive genome of loblolly pine using haploid DNA and novel assembly strategies". Genome Biology. 15 (3): R59. doi:10.1186/gb-2014-15-3-r59. PMC 4053751. PMID 24647006.
- ^ Pellicer J, Fay MF, Leitch IJ (15 September 2010). "The largest eukaryotic genome of them all?". Botanical Journal of the Linnean Society. 164 (1): 10–15. doi:10.1111/j.1095-8339.2010.01072.x.
- ^ Lang D, Zimmer AD, Rensing SA, Reski R (October 2008). "Exploring plant biodiversity: the Physcomitrella genome and beyond". Trends in Plant Science. 13 (10): 542–49. doi:10.1016/j.tplants.2008.07.002. PMID 18762443.
- ^ "Saccharomyces Genome Database". Yeastgenome.org. Retrieved 27 January 2011.
- ^ Galagan JE, Calvo SE, Cuomo C, Ma LJ, Wortman JR, Batzoglou S, et al. (December 2005). "Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae". Nature. 438 (7071): 1105–15. Bibcode:2005Natur.438.1105G. doi:10.1038/nature04341. PMID 16372000.
- ^ Leroy S, Bouamer S, Morand S, Fargette M (2007). "Genome size of plant-parasitic nematodes". Nematology. 9 (3): 449–50. doi:10.1163/156854107781352089.
- ^ Gregory TR (2005). "Animal Genome Size Database". Gregory, T.R. (2016). Animal Genome Size Database.
- ^ The C. elegans Sequencing Consortium (December 1998). "Genome sequence of the nematode C. elegans: a platform for investigating biology". Science. 282 (5396): 2012–18. Bibcode:1998Sci...282.2012.. doi:10.1126/science.282.5396.2012. PMID 9851916. S2CID 16873716.
- ^ Kelley, Joanna L.; Peyton, Justin T.; Fiston-Lavier, Anna-Sophie; Teets, Nicholas M.; Yee, Muh-Ching; Johnston, J. Spencer; Bustamante, Carlos D.; Lee, Richard E.; Denlinger, David L. (12 August 2014). "Compact genome of the Antarctic midge is likely an adaptation to an extreme environment". Nature Communications. 5: 4611. Bibcode:2014NatCo...5.4611K. doi:10.1038/ncomms5611. ISSN 2041-1723. PMC 4164542. PMID 25118180.
- ^ Ellis LL, Huang W, Quinn AM, Ahuja A, Alfrejd B, Gomez FE, Hjelmen CE, Moore KL, Mackay TF, Johnston JS, Tarone AM (July 2014). "Intrapopulation genome size variation in D. melanogaster reflects life history variation and plasticity". PLOS Genetics. 10 (7): e1004522. doi:10.1371/journal.pgen.1004522. PMC 4109859. PMID 25057905.
- ^ Honeybee Genome Sequencing Consortium (October 2006). "Insights into social insects from the genome of the honeybee Apis mellifera". Nature. 443 (7114): 931–49. Bibcode:2006Natur.443..931T. doi:10.1038/nature05260. PMC 2048586. PMID 17073008.
- ^ The International Silkworm Genome (December 2008). "The genome of a lepidopteran model insect, the silkworm Bombyx mori". Insect Biochemistry and Molecular Biology. 38 (12): 1036–45. doi:10.1016/j.ibmb.2008.11.004. PMID 19121390.
- ^ Wurm Y, Wang J, Riba-Grognuz O, Corona M, Nygaard S, Hunt BG, et al. (April 2011). "The genome of the fire ant Solenopsis invicta". Proceedings of the National Academy of Sciences of the United States of America. 108 (14): 5679–84. Bibcode:2011PNAS..108.5679W. doi:10.1073/pnas.1009690108. PMC 3078418. PMID 21282665.
- ^ Shao, Changwei; Sun, Shuai; Liu, Kaiqiang; Wang, Jiahao; Li, Shuo; Liu, Qun; Deagle, Bruce E.; Seim, Inge; Biscontin, Alberto; Wang, Qian; Liu, Xin; Kawaguchi, So; Liu, Yalin; Jarman, Simon; Wang, Yue (16 March 2023). "The enormous repetitive Antarctic krill genome reveals environmental adaptations and population insights". Cell Glish. 186 (6): 1279–1294.e19. doi:10.1016/j.cell.2023.02.005. hdl:11577/3472081. ISSN 0092-8674. PMID 36868220. S2CID 257286259.
- ^ "Junk DNA Deforms Salamander Bodies". Scientific American. February 2022. Archived from the original on 22 May 2023.
- ^ A bird-like genome from a frog - PNAS
- ^ Ecological factors and parity mode correlate with genome size variation in squamate reptiles
- ^ Evolution of the Largest Mammalian Genome
- ^ Church DM, Goodstadt L, Hillier LW, Zody MC, Goldstein S, She X, et al. (May 2009). Roberts RJ (ed.). "Lineage-specific biology revealed by a finished genome assembly of the mouse". PLOS Biology. 7 (5): e1000112. doi:10.1371/journal.pbio.1000112. PMC 2680341. PMID 19468303.
- ^ "Pan paniscus (pygmy chimpanzee)". nih.gov. Retrieved 30 June 2016.
- ^ Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, et al. (February 2001). "The sequence of the human genome". Science. 291 (5507): 1304–51. Bibcode:2001Sci...291.1304V. doi:10.1126/science.1058040. PMID 11181995.
- ^ International Chicken Genome Sequencing Consortium (December 2004). "Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution". Nature. 432 (7018): 695–716. Bibcode:2004Natur.432..695C. doi:10.1038/nature03154. ISSN 0028-0836. PMID 15592404.
- ^ Roest Crollius H, Jaillon O, Dasilva C, Ozouf-Costaz C, Fizames C, Fischer C, Bouneau L, Billault A, Quetier F, Saurin W, Bernot A, Weissenbach J (July 2000). "Characterization and repeat analysis of the compact genome of the freshwater pufferfish Tetraodon nigroviridis". Genome Research. 10 (7): 939–49. doi:10.1101/gr.10.7.939. PMC 310905. PMID 10899143.
- ^ Jaillon O, Aury JM, Brunet F, Petit JL, Stange-Thomann N, Mauceli E, et al. (October 2004). "Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype". Nature. 431 (7011): 946–57. Bibcode:2004Natur.431..946J. doi:10.1038/nature03025. PMID 15496914.
- ^ "Tetraodon Project Information". Archived from the original on 26 September 2012. Retrieved 17 October 2012.
- ^ Leitch, I J (August 2007). "Genome sizes through the ages". Heredity. 99 (2): 121–122. doi:10.1038/sj.hdy.6800981. ISSN 0018-067X.
- ^ "Genome size and chromosome number database we - Various - BNID 100597". bionumbers.hms.harvard.edu. Retrieved 17 August 2024.
- ^ "Animal Genome Size Database:: Statistics". www.genomesize.com. Retrieved 17 August 2024.
- ^ Zimmer, Carl (31 May 2024). "Scientists Find the Largest Known Genome Inside a Small Plant - A fern from a Pacific island carries 50 times as much DNA as humans do". The New York Times. Archived from the original on 31 May 2024. Retrieved 1 June 2024.
- ^ Martincorena I, Campbell PJ (September 2015). "Somatic mutation in cancer and normal cells". Science. 349 (6255): 1483–89. Bibcode:2015Sci...349.1483M. doi:10.1126/science.aab4082. PMID 26404825. S2CID 13945473.
- ^ Hajkova P, Jeffries SJ, Lee C, Miller N, Jackson SP, Surani MA (July 2010). "Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway". Science. 329 (5987): 78–82. Bibcode:2010Sci...329...78H. doi:10.1126/science.1187945. PMC 3863715. PMID 20595612.
- ^ Hackett JA, Sengupta R, Zylicz JJ, Murakami K, Lee C, Down TA, Surani MA (January 2013). "Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine". Science. 339 (6118): 448–52. Bibcode:2013Sci...339..448H. doi:10.1126/science.1229277. PMC 3847602. PMID 23223451.
- ^ "Gattaca (movie)". Rotten Tomatoes. 24 October 1997.
Further reading
[edit]- Benfey P, Protopapas AD (2004). Essentials of Genomics. Prentice Hall.
- Brown TA (2002). Genomes 2. Oxford: Bios Scientific Publishers. ISBN 978-1-85996-029-5.
- Gibson G, Muse SV (2004). A Primer of Genome Science (Second ed.). Sunderland, Mass: Sinauer Assoc. ISBN 978-0-87893-234-4.
- Gregory TR (2005). The Evolution of the Genome. Elsevier. ISBN 978-0-12-301463-4.
- Reece RJ (2004). Analysis of Genes and Genomes. Chichester: John Wiley & Sons. ISBN 978-0-470-84379-6.
- Saccone C, Pesole G (2003). Handbook of Comparative Genomics. Chichester: John Wiley & Sons. ISBN 978-0-471-39128-9.
- Werner E (December 2003). "In silico multicellular systems biology and minimal genomes". Drug Discovery Today. 8 (24): 1121–27. doi:10.1016/S1359-6446(03)02918-0. PMID 14678738.
External links
[edit]- UCSC Genome Browser – view the genome and annotations for more than 80 organisms.
- genomecenter.howard.edu (archived 9 August 2013)
- Build a DNA Molecule (archived 9 June 2010)
- Some comparative genome sizes
- DNA Interactive: The History of DNA Science
- DNA From The Beginning
- All About The Human Genome Project—from Genome.gov
- Animal genome size database
- Plant genome size database (archived 1 September 2005)
- GOLD:Genomes OnLine Database
- The Genome News Network
- NCBI Entrez Genome Project database
- NCBI Genome Primer
- GeneCards—an integrated database of human genes
- BBC News – Final genome 'chapter' published
- IMG (The Integrated Microbial Genomes system)—for genome analysis by the DOE-JGI
- GeKnome Technologies Next-Gen Sequencing Data Analysis—next-generation sequencing data analysis for Illumina and 454 Service from GeKnome Technologies (archived 3 March 2012)
